Documenti di Didattica
Documenti di Professioni
Documenti di Cultura
Allele Frequencies of The New European Standard Set (ESS) Loci Plus SE33 Locus in A Population From The Republic of Macedonia
Caricato da
conteconteDescrizione originale:
Titolo originale
Copyright
Formati disponibili
Condividi questo documento
Condividi o incorpora il documento
Hai trovato utile questo documento?
Questo contenuto è inappropriato?
Segnala questo documentoCopyright:
Formati disponibili
Allele Frequencies of The New European Standard Set (ESS) Loci Plus SE33 Locus in A Population From The Republic of Macedonia
Caricato da
conteconteCopyright:
Formati disponibili
G Model
FSIGEN-770; No. of Pages 3
Forensic Science International: Genetics xxx (2011) xxxxxx
Contents lists available at ScienceDirect
Forensic Science International: Genetics
journal homepage: www.elsevier.com/locate/fsig
Letter to the Editor
Allele frequencies of the new European Standard Set (ESS) loci plus SE33 locus in a population from the Republic of Macedonia Dear Editor, The highly polymorphic short tandem repeat (STR) loci in the human genome are widely used for forensic and paternity testing as well as for population genetic studies. Introducing set of STR loci as observed markers made marvels progress in this eld of forensic science [1,2]. Nowadays, in response to the ENFSI and EDNAP groups call for new STR multiplexes for Europe, Promega developed a suite of four new DNA proling kits. One of them is PowerPlex ESI 17 that we have used in this study. In our previous population studies of Macedonian human population, we have used 15 STR loci included in the AmpFlSTR1Identiler1 [3] and 17 Y-chromosomal short tandem repeats loci incorporated in the AmpFistr1YlerTM PCR Amplication Kit (data submitted to the FSI Genetics). All obtained results were included in Macedonian referent database. In order of future development of this database we have decide to analyze 6 additional new STR loci (D22S1045, D1S1656, D10S1248, D2S441, D12S391, and SE33). Some of these loci were described even 15 years ago [4,5]. This study is based on an observation of representative sample of Macedonia human population. A total of 205 unrelated individuals of which 105 males and 100 females, from different regions of Macedonia, participated in this study. All tested individuals have been voluntary donors appropriately informed about the research. Qiagen MicroTM Tissue Kit was used for DNA extraction from whole blood, Quantiler1 assay for quantication and PowerPlex ESI 171 System for amplication and detection (D18S51, D21S11, TH01, D3S1358, Amelogenin, D16S539, D2S1338, D1S1656, D10S1248, FGA, D8S1179, vWA, D22S1045, D19S433, D12S391, D2S441 and SE33). The total volume of each reaction was 12.5 ml. The PCR amplications have been carried out in ABI 9700 Thermal Cycler according to the manufacturers recommendations. Electrophoresis of the amplication products was performed on an ABI 310 PRISM. Numerical allele designations of the proles were obtained by processing with Gene Mapper1 IDv 3.2 Software. Deviation from HardyWeinberg equilibrium, observed and expected heterozygosity, power of discrimination and power of exclusion were calculated. Also, we have compared Macedonian data with data obtained from other neighboring populations with available data for observed loci. Deviation from HardyWeinberg equilibrium [6], observed and expected heterozygosity [7], exact test of population differentiation [8] and pairwise Fst [9] were calculated within Powermarker v3.25 [10], since power of discrimination and power of exclusion within Microsoft1 Excel workbook templatePowerStats [11]. Allele frequencies and all calculated forensic parameters are shown in Supplementary Table 1. All data are available on http:// www.ingeb.ba/rebida/ESI17/macedonia.html. No statistically signicant deviation (p < 0.05) from HardyWeinberg equilibrium was found except for D1S1656 and SE33 loci (Table 1). Heterozygosity excess has been detected for SE33 locus. Allele frequencies of observed six loci within Macedonian population have been compared with data obtained from geographically neighboring Southeast European populations such as Serbia and Bosnia Herzegovina (our unpublished data). After Bonferronis correction, statistical signicance was considered as
Table 1 The distribution of allele frequencies and calculated forensic parameters for the Macedonian population over 5 new ESS loci and SE33 locus (n = 205). Allele 2n 1 6.3 9 10 11 11.3 12 12.1 12.3 13 13.2 14 14.2 15 15.3 16 16.1 16.3 17 D1S1656 410 0.1146 0.1683 0.0683 0.0780 0.1244 0.0463 0.1366 0.0390 0.0341 D2S441 410 0.0024 0.1268 0.3365 0.0951 0.0415 0.0039 0.0024 0.0390 0.3073 0.0439 0.0024 D10S1248 410 0.0268 0.2341 0.3293 0.1780 0.1829 0.0439 D12S391 410 0.0049 0.0024 0.0512 0.0317 0.1000 SE33 410 0.0024 0.0024 0.0254 0.0122 0.0073 0.0098 0.0073 0.0463 0.0366 0.0024 0.0659 D22S1045 410 0.0024 0.1366 0.0122 0.0537 0.3634 0.3220 0.0951
1872-4973/$ see front matter 2011 Elsevier Ireland Ltd. All rights reserved. doi:10.1016/j.fsigen.2011.07.004
Please cite this article in press as: Z. Jakovski, et al., Allele frequencies of the new European Standard Set (ESS) loci plus SE33 locus in a population from the Republic of Macedonia, Book review. Forensic Sci. Int. Genet. (2011), doi:10.1016/j.fsigen.2011.07.004
G Model
FSIGEN-770; No. of Pages 3
2 Table 1 (Continued ) Allele 2n 17.2 17.3 18 19 19.2 19.3 20 20.2 20.3 21 21.2 22 22.2 23 24 24.1 24.2 25 25.2 26 27 27.2 28 28.2 29 29.2 30.2 31.2 32.2 33 33.2 34.2 35 36 37.2 PD PE P H(ex) H(ob) D1S1656 410 0.0902 0.0146 0.0024 0.0098 0.973 0.791 0.0223 0.8950 0.8976 D2S441 410 0.929 0.506 0.1076 0.7636 0.6976 D10S1248 410 0.0024 0.0024 0.902 0.599 0.2502 0.7689 0.8000 D12S391 410 0.0024 0.1732 0.1171 0.0049 0.0902 0.0024 0.1317 0.1146 0.0707 0.0463 0.0293 0.0024 0.0024 0.976 0.721 0.0582 0.8955 0.8634 SE33 410 0.0049 0.0756 0.1098 0.0049 0.0537 0.0122 0.0122 0.0122 0.0073 0.0366 0.0024 0.0024 0.0390 0.0024 0.0244 0.0024 0.0756 0.0024 0.0512 0.0024 0.0610 0.0488 0.0146 0.0220 0.0024 0.0073 0.0024 0.0073 0.0024 0.0024 0.989 0.810 0.0029 0.9473 0.9073 D22S1045 410 0.0073 0.0073 0.878 0.579 0.3163 0.7334 0.7854 Letter to the Editor / Forensic Science International: Genetics xxx (2011) xxxxxx
PD, power of discrimination; PE, power of exclusion; P, HardyWeinberg equilibrium, exact test; H(ob), observed heterozygosity; H(ex), expected heterozygosity.
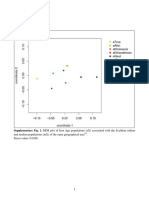
p < 0.015. Statistically signicant differences of allele frequencies were noticed for D12S391 locus (p = 0.0019) between Macedonian and Serbian population, as well as between Macedonian and Bosnia Herzegovina population (p = 0.0005). Pairwise Fst, calculated according to the results observed over these 6 loci, shows no signicant genetic difference among Macedonian and Serbian population (0.14%), as well as among Macedonian and Bosnia Herzegovina population (0.15%). According to our preliminary results, no disconcordance between PowerPlex ESI 17 and Identiler or PowerPlex 16 was recorded. Detailed concordance study, which is going to include almost thousand samples from this part of Europe, is ongoing. In the course of the elaboration of this population study, we adhered to the new guidelines published in this journal, wherein we would like to point out that we accept the requirements pertaining to the publication of this type of papers [12], as well a ISFG recommendations [13]. Also we follow recommendations of the European Council resolution (9192/01) calls upon EU countries to use the European Standard Set (ESS) as a minimum to enable international comparison of DNA proles as well as the paper for the evolution of DNA databases and recommendations [14]. Acknowledgment We would like to thank to the Promega Corporation for supporting this study.
References
[1] P. Gill, C.P. Kimpton, A. Urquhart, et al., Automated short tandem repeat (STR) analysis in forensic caseworka strategy for the future, Electrophoresis 16 (1995) 15431552. [2] J.A. Thomson, V. Pilotti, P. Stevens, K.L. Ayres, P.G. Debenham, Validation of short tandem repeat analysis for the investigation of cases of disputed paternity, Forensic Sci. Int. 100 (1998) 116. [3] Z. Jakovski, K. Nikolova, I. Furac, M. Masic, B. Janeska, M. Kubat, Allele frequencies for 15 STR loci in a population from the Republic of Macedonia, Int. J. Legal Med. 120 (2006) 5355. [4] M.V. Lareu, C. Pestoni, M. Schurenkamp, S. Rand, B. Brinkmann, A. Carracedo, A highly variable STR at the D12S391 locus, Int. J. Legal Med. 109 (3) (1996) 134138. [5] M.V. Lareu, S. Barral, A. Salas, C. Pestoni, A. Carracedo, Sequence variation of a hypervariable short tandem repeat at the D1S1656 locus, Int. J. Legal Med. 111 (5) (1998) 244247. [6] S.W. Guo, E.A. Thompson, Performing the exact test of HardyWeinberg proportion for multiple alleles, Biometrics 48 (1992) 361372. [7] M. Nei, Molecular Evolutionary Genetics, Columbia University Press, New York, 1987. [8] M. Raymond, F. Rousset, An exact test for population differentiation, Evolution 49 (1995) 12801283. [9] B.S. Weir, Genetic Data Analysis II: Methods for Discrete Population Genetic Data, Sinauer Assoc., Inc., Sunderland, MA, 1996. [10] L. Kejun, V.M. Spencer, PowerMarker: an integrated analysis environment for genetic marker analysis, Bioinformatics 21 (2005) 21282129. [11] A. Tereba, Tools for analysis of population statistics, Proles DNA 3 (1999) 1416. [12] A. Carracedo, J.M. Butler, L. Gusmao, W. Parson, L. Roewer, P.M. Schneider, Publication of population data for forensic purposes, Forensic Sci. Int. Genet. 4 (2010) 145147. [13] B. Olaisen, W. Bar, B. Brinkmann, B. Budowle, A. Carracedo, P. Gill, P. Lincoln, W.R. Mayr, S. Vox Sang Rand, DNA recommendations 1997 of the International Society for Forensic Genetics, Vox Sang. 74 (1) (1998) 6163.
Please cite this article in press as: Z. Jakovski, et al., Allele frequencies of the new European Standard Set (ESS) loci plus SE33 locus in a population from the Republic of Macedonia, Book review. Forensic Sci. Int. Genet. (2011), doi:10.1016/j.fsigen.2011.07.004
G Model
FSIGEN-770; No. of Pages 3
Letter to the Editor / Forensic Science International: Genetics xxx (2011) xxxxxx [14] P. Gill, L. Fereday, N. Morling, P.M. Schneider, The evolution of DNA databases recommendations for new European STR loci, Forensic Sci. Int. 156 (2006) 242244. 3
Zlatko Jakovski* Ksenija Nikolova Renata Jankova-Ajanovska Biljana Janeska Institute of Forensic Medicine and Criminology, School of Medicine, University Ss. Cyril and Methodious, Skopje, Macedonia Naris Pojskic Institute for Genetic Engineering and Biotechnology, Sarajevo, Bosnia and Herzegovina
Damir Marjanovicab Institute for Genetic Engineering and Biotechnology, Sarajevo, Bosnia and Herzegovina b Genos, Planinska 1, 10000 Zagreb, Croatia
*Corresponding
author. Tel.: +389 2 3 177 044; fax: +389 2 1 178 831
E-mail address: zlatedr@yahoo.com (Z. Jakovski). 24 December 2010
Please cite this article in press as: Z. Jakovski, et al., Allele frequencies of the new European Standard Set (ESS) loci plus SE33 locus in a population from the Republic of Macedonia, Book review. Forensic Sci. Int. Genet. (2011), doi:10.1016/j.fsigen.2011.07.004
Potrebbero piacerti anche
- The Yellow House: A Memoir (2019 National Book Award Winner)Da EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Valutazione: 4 su 5 stelle4/5 (98)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceDa EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceValutazione: 4 su 5 stelle4/5 (895)
- Book - of - Abstracts - Isabs - 2015 - 2 55Documento1 paginaBook - of - Abstracts - Isabs - 2015 - 2 55conteconteNessuna valutazione finora
- Paternal and Maternal Lineages in The BalkansDocumento29 paginePaternal and Maternal Lineages in The BalkansdfwefeNessuna valutazione finora
- Ban JaniDocumento1 paginaBan JaniconteconteNessuna valutazione finora
- Paternal and Maternal Lineages in The BalkansDocumento29 paginePaternal and Maternal Lineages in The BalkansdfwefeNessuna valutazione finora
- Godisnjak 13Documento14 pagineGodisnjak 13conteconteNessuna valutazione finora
- Ncomms14615 s1 PDFDocumento82 pagineNcomms14615 s1 PDFconteconteNessuna valutazione finora
- Ivan LuciusDocumento1 paginaIvan LuciusconteconteNessuna valutazione finora
- Sanok DistrictDocumento55 pagineSanok DistrictconteconteNessuna valutazione finora
- Allele Frequencies 15 AmpFISTR - Vojvodina, Serbia, MontenegroDocumento3 pagineAllele Frequencies 15 AmpFISTR - Vojvodina, Serbia, MontenegroconteconteNessuna valutazione finora
- Genetic Data From Y Chromosome STR and SNP Loci in Ukrainian PopulationDocumento4 pagineGenetic Data From Y Chromosome STR and SNP Loci in Ukrainian PopulationconteconteNessuna valutazione finora
- South East RomaniaDocumento6 pagineSouth East RomaniaBranislav Dinic100% (1)
- Y-STR Variation in Albanian PopulationsDocumento8 pagineY-STR Variation in Albanian PopulationsconteconteNessuna valutazione finora
- Medhist00142 0057Documento8 pagineMedhist00142 0057conteconteNessuna valutazione finora
- Directive 2003-54-EC of EU Parlament of 26.june 2003Documento19 pagineDirective 2003-54-EC of EU Parlament of 26.june 2003conteconteNessuna valutazione finora
- Adeline Paulina IrbyDocumento7 pagineAdeline Paulina IrbyJosh Irby100% (3)
- Jugoistočna Evropa 2005Documento12 pagineJugoistočna Evropa 2005conteconteNessuna valutazione finora
- Genetic Polymorphism in A Serbian PopulationDocumento2 pagineGenetic Polymorphism in A Serbian PopulationconteconteNessuna valutazione finora
- Allele Frequencies 15 AmpFISTR - Vojvodina, Serbia, MontenegroDocumento3 pagineAllele Frequencies 15 AmpFISTR - Vojvodina, Serbia, MontenegroconteconteNessuna valutazione finora
- Genetic Polymorphism in A Serbian PopulationDocumento2 pagineGenetic Polymorphism in A Serbian PopulationconteconteNessuna valutazione finora
- Izvestaj o Medijima Preciscen Eng.Documento44 pagineIzvestaj o Medijima Preciscen Eng.siriusistNessuna valutazione finora
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeDa EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeValutazione: 4 su 5 stelle4/5 (5794)
- The Little Book of Hygge: Danish Secrets to Happy LivingDa EverandThe Little Book of Hygge: Danish Secrets to Happy LivingValutazione: 3.5 su 5 stelle3.5/5 (399)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaDa EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaValutazione: 4.5 su 5 stelle4.5/5 (266)
- Shoe Dog: A Memoir by the Creator of NikeDa EverandShoe Dog: A Memoir by the Creator of NikeValutazione: 4.5 su 5 stelle4.5/5 (537)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureDa EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureValutazione: 4.5 su 5 stelle4.5/5 (474)
- Never Split the Difference: Negotiating As If Your Life Depended On ItDa EverandNever Split the Difference: Negotiating As If Your Life Depended On ItValutazione: 4.5 su 5 stelle4.5/5 (838)
- Grit: The Power of Passion and PerseveranceDa EverandGrit: The Power of Passion and PerseveranceValutazione: 4 su 5 stelle4/5 (588)
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryDa EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryValutazione: 3.5 su 5 stelle3.5/5 (231)
- The Emperor of All Maladies: A Biography of CancerDa EverandThe Emperor of All Maladies: A Biography of CancerValutazione: 4.5 su 5 stelle4.5/5 (271)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyDa EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyValutazione: 3.5 su 5 stelle3.5/5 (2259)
- On Fire: The (Burning) Case for a Green New DealDa EverandOn Fire: The (Burning) Case for a Green New DealValutazione: 4 su 5 stelle4/5 (73)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersDa EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersValutazione: 4.5 su 5 stelle4.5/5 (344)
- Team of Rivals: The Political Genius of Abraham LincolnDa EverandTeam of Rivals: The Political Genius of Abraham LincolnValutazione: 4.5 su 5 stelle4.5/5 (234)
- The Unwinding: An Inner History of the New AmericaDa EverandThe Unwinding: An Inner History of the New AmericaValutazione: 4 su 5 stelle4/5 (45)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreDa EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreValutazione: 4 su 5 stelle4/5 (1090)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)Da EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Valutazione: 4.5 su 5 stelle4.5/5 (121)
- Her Body and Other Parties: StoriesDa EverandHer Body and Other Parties: StoriesValutazione: 4 su 5 stelle4/5 (821)
- Discromatie Test 38Documento13 pagineDiscromatie Test 38nelutza_popNessuna valutazione finora
- Test Bank For Molecular Biology Principles and Practice 1st Edition Michael M CoxDocumento9 pagineTest Bank For Molecular Biology Principles and Practice 1st Edition Michael M CoxAnthonyRogersydtfp100% (65)
- Intermediate Genetics ProblemsDocumento4 pagineIntermediate Genetics Problemsandrea lusinguNessuna valutazione finora
- Making A Model of DNA InstructionsDocumento9 pagineMaking A Model of DNA Instructionsapi-256992527Nessuna valutazione finora
- Cer Chart For Who Killed Carleton CometDocumento2 pagineCer Chart For Who Killed Carleton Cometapi-336683878Nessuna valutazione finora
- Lecture 1 - Mendelian Laws of InheritanceDocumento31 pagineLecture 1 - Mendelian Laws of InheritancekibzwanjikuNessuna valutazione finora
- Advance Blast Rani Anak Mat 212111Documento3 pagineAdvance Blast Rani Anak Mat 212111Rani anak matNessuna valutazione finora
- Mitosis and Meiosis: Bio Lab-131 Session-Fall 2021 Section-001Documento27 pagineMitosis and Meiosis: Bio Lab-131 Session-Fall 2021 Section-001Satya NadellaNessuna valutazione finora
- Molecular Diagnosis of Diseases and Parental TestingDocumento16 pagineMolecular Diagnosis of Diseases and Parental Testingmuhammad zakriaNessuna valutazione finora
- c13 Microbiology Tortora TestbankDocumento17 paginec13 Microbiology Tortora Testbankwhitewave25Nessuna valutazione finora
- The Concept of Society - The Components and Their InterrelationsDocumento18 pagineThe Concept of Society - The Components and Their InterrelationsSofia LatorreNessuna valutazione finora
- GeneticsDocumento34 pagineGeneticsJuan Carlos Dela CruzNessuna valutazione finora
- Bio BibliographyDocumento5 pagineBio BibliographyJamieNessuna valutazione finora
- Week 3c - Phylogenetic - Tree - ConstructionMai PDFDocumento19 pagineWeek 3c - Phylogenetic - Tree - ConstructionMai PDFSunilNessuna valutazione finora
- Walter Gilbert RNA WorldDocumento1 paginaWalter Gilbert RNA WorldARGHA MANNANessuna valutazione finora
- Ms Campbell Protein Synthesis Practice Questions Regents LeDocumento6 pagineMs Campbell Protein Synthesis Practice Questions Regents Lekim del mundoNessuna valutazione finora
- Biochemistry Final ExamDocumento2 pagineBiochemistry Final ExamJesson BelenNessuna valutazione finora
- Molecular Basis of Inheritance, Class 12, Cbse NotesDocumento33 pagineMolecular Basis of Inheritance, Class 12, Cbse NotesSubho Bhattacharya50% (4)
- BIOL 311 Mapping ReportDocumento8 pagineBIOL 311 Mapping ReportJermaine I-Inowculate Parker100% (1)
- Journal of Plant PhysiologyDocumento4 pagineJournal of Plant PhysiologygillNessuna valutazione finora
- MitoDocumento30 pagineMitoabeer adelNessuna valutazione finora
- Cmap Science 9Documento3 pagineCmap Science 9Zharina Ann EstavilloNessuna valutazione finora
- Bio (In Focus Year 12)Documento636 pagineBio (In Focus Year 12)isra el100% (1)
- Biology Form 5: Chapter 6 (Variation)Documento5 pagineBiology Form 5: Chapter 6 (Variation)Gerard Selvaraj100% (5)
- Cell Final ExamDocumento8 pagineCell Final ExamJorge GomezNessuna valutazione finora
- Gene PredictionDocumento50 pagineGene Predictionsaeedahmad901Nessuna valutazione finora
- Heredity and GeneticsDocumento20 pagineHeredity and GeneticssrichardequipNessuna valutazione finora
- Genetic Calculator (Budgerigar) Help File: © 2016 K YorkeDocumento22 pagineGenetic Calculator (Budgerigar) Help File: © 2016 K YorkeMarlindaNessuna valutazione finora
- Sgi Evolution Ss 11 1Documento6 pagineSgi Evolution Ss 11 1api-329922192Nessuna valutazione finora
- Fears of A Brave New WorldDocumento6 pagineFears of A Brave New WorldPier Paolo Dal MonteNessuna valutazione finora