Documenti di Didattica
Documenti di Professioni
Documenti di Cultura
1.7 Sanger Sequencing
Caricato da
Utsav Ullas0 valutazioniIl 0% ha trovato utile questo documento (0 voti)
21 visualizzazioni20 pagineSanger sequencing is a method for DNA sequencing developed by Frederick Sanger in 1977. It involves using DNA polymerase to selectively incorporate chain-terminating dideoxynucleotides during in vitro DNA replication, generating DNA fragments of different lengths that can be separated by gel electrophoresis. The order of the bases is read from the gel according to the fragment lengths, allowing determination of the DNA sequence. Sanger sequencing revolutionized DNA sequencing and was the primary method of DNA sequencing until the development of next-generation sequencing technologies.
Descrizione originale:
Sanger sequencing a presentational idea.
Copyright
© © All Rights Reserved
Formati disponibili
PDF, TXT o leggi online da Scribd
Condividi questo documento
Condividi o incorpora il documento
Hai trovato utile questo documento?
Questo contenuto è inappropriato?
Segnala questo documentoSanger sequencing is a method for DNA sequencing developed by Frederick Sanger in 1977. It involves using DNA polymerase to selectively incorporate chain-terminating dideoxynucleotides during in vitro DNA replication, generating DNA fragments of different lengths that can be separated by gel electrophoresis. The order of the bases is read from the gel according to the fragment lengths, allowing determination of the DNA sequence. Sanger sequencing revolutionized DNA sequencing and was the primary method of DNA sequencing until the development of next-generation sequencing technologies.
Copyright:
© All Rights Reserved
Formati disponibili
Scarica in formato PDF, TXT o leggi online su Scribd
0 valutazioniIl 0% ha trovato utile questo documento (0 voti)
21 visualizzazioni20 pagine1.7 Sanger Sequencing
Caricato da
Utsav UllasSanger sequencing is a method for DNA sequencing developed by Frederick Sanger in 1977. It involves using DNA polymerase to selectively incorporate chain-terminating dideoxynucleotides during in vitro DNA replication, generating DNA fragments of different lengths that can be separated by gel electrophoresis. The order of the bases is read from the gel according to the fragment lengths, allowing determination of the DNA sequence. Sanger sequencing revolutionized DNA sequencing and was the primary method of DNA sequencing until the development of next-generation sequencing technologies.
Copyright:
© All Rights Reserved
Formati disponibili
Scarica in formato PDF, TXT o leggi online su Scribd
Sei sulla pagina 1di 20
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
1
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
2
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
3
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
4
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
5
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
6
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
7
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
8
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
9
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
10
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
11
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
12
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
13
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
14
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
15
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
16
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
17
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
18
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
19
Genomic Technologies in Clinical Diagnostics
Sanger Sequencing
Video animation provided by Wellcome Trust
20
Potrebbero piacerti anche
- HMX CourseOutlines Corp PGEN SEQ v03Documento1 paginaHMX CourseOutlines Corp PGEN SEQ v03Saitama LuffyNessuna valutazione finora
- ADVIA Centaur CP - SiemensDocumento2 pagineADVIA Centaur CP - SiemensYann Christian HIENNessuna valutazione finora
- 121 GD PrenatalScreeningGuide Jan21Documento3 pagine121 GD PrenatalScreeningGuide Jan21Jyotsnna GuptaaNessuna valutazione finora
- Flyer-Tpoo1-06-En HJK LongDocumento4 pagineFlyer-Tpoo1-06-En HJK LongG.330Nessuna valutazione finora
- Thermo Scientific Forma High-Performance Laboratory Refrigerators and FreezersDocumento22 pagineThermo Scientific Forma High-Performance Laboratory Refrigerators and FreezerswhoisvicksNessuna valutazione finora
- Thermo Scientific Product Catalog: Drugs of Abuse Testing, Drug Monitoring, Instrumentation, and Quality Control ProductsDocumento41 pagineThermo Scientific Product Catalog: Drugs of Abuse Testing, Drug Monitoring, Instrumentation, and Quality Control ProductsAntónio FreitasNessuna valutazione finora
- Diagnostic Assays & Instruments: Gold Standard Diagnostics Europe 2022 International CatalogueDocumento28 pagineDiagnostic Assays & Instruments: Gold Standard Diagnostics Europe 2022 International CatalogueMentor KurshumliuNessuna valutazione finora
- 2midterm 8 - Diagnostic Virology TRANSDocumento2 pagine2midterm 8 - Diagnostic Virology TRANSRobee Camille Desabelle-SumatraNessuna valutazione finora
- Simple, Point-Of-Care Urinalysis With Enhanced Clinical InformationDocumento4 pagineSimple, Point-Of-Care Urinalysis With Enhanced Clinical InformationZakaria Al SharabiNessuna valutazione finora
- CLP Tech Guide Molecular Diagnostic InstrumentsDocumento2 pagineCLP Tech Guide Molecular Diagnostic InstrumentsKian KianNessuna valutazione finora
- PG2403 PJT7091 COL34261 GeneArt ResourceGuide Global FWRDocumento36 paginePG2403 PJT7091 COL34261 GeneArt ResourceGuide Global FWRpastorrlc8852Nessuna valutazione finora
- HindiDocumento5 pagineHindiharshita nairNessuna valutazione finora
- 1-Localizing Anomalies From Weakly Labeled VideosDocumento10 pagine1-Localizing Anomalies From Weakly Labeled VideosEsraa AlaaNessuna valutazione finora
- Brosur Clinitek Novus BrochureDocumento2 pagineBrosur Clinitek Novus BrochureFarly AugusNessuna valutazione finora
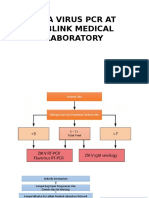
- Zika Virus PCR at Lablink Medical LaboratoryDocumento13 pagineZika Virus PCR at Lablink Medical LaboratoryAmrit SoniaNessuna valutazione finora
- Sample Report CentoNIPT Positive-Findings 20230428Documento3 pagineSample Report CentoNIPT Positive-Findings 20230428Zbant Liviu GabrielNessuna valutazione finora
- V2.2023 Concert Genetic Testing - Non-Invasive Prenatal Screening (NIPS)Documento13 pagineV2.2023 Concert Genetic Testing - Non-Invasive Prenatal Screening (NIPS)IOAN STOICANessuna valutazione finora
- Lab TestDocumento13 pagineLab TestAnonymous VsPjwVNessuna valutazione finora
- Debatingdenim 48x36Documento1 paginaDebatingdenim 48x36eng213810140Nessuna valutazione finora
- Abbott HPV RealTime AssayDocumento2 pagineAbbott HPV RealTime AssayIbrehimaNessuna valutazione finora
- Akarslan2008 PDFDocumento6 pagineAkarslan2008 PDFLaura ImreNessuna valutazione finora
- S - Diseases - PDF (Google Scholar)Documento3 pagineS - Diseases - PDF (Google Scholar)Dandi SeptianNessuna valutazione finora
- GP Letak A4 HBV IVDD EN 5 2023Documento2 pagineGP Letak A4 HBV IVDD EN 5 2023Fatima VessaliusNessuna valutazione finora
- Fast Track Diagnostics Assays Brochure 0917 FINAL 1800000004342593Documento6 pagineFast Track Diagnostics Assays Brochure 0917 FINAL 1800000004342593Svasthya ManagerNessuna valutazione finora
- Thymovac - Product IntroductionDocumento20 pagineThymovac - Product Introductionnquockhanh1998Nessuna valutazione finora
- Corporate Profile 2021NABLDocumento27 pagineCorporate Profile 2021NABLSuraj ChauguleNessuna valutazione finora
- 21 HPV GenoArray Diagnostic KitDocumento8 pagine21 HPV GenoArray Diagnostic KitYosinee PatrungsiNessuna valutazione finora
- Brochure-Real-time PCR Detection Kit For Monkeypox VirusDocumento2 pagineBrochure-Real-time PCR Detection Kit For Monkeypox VirusMouhamadou Habibi Keys TraoreNessuna valutazione finora
- QIAseq Targeted DNA Pro Workshop - Panel PresentationDocumento21 pagineQIAseq Targeted DNA Pro Workshop - Panel PresentationRini HafzariNessuna valutazione finora
- Ncm106 Course OutlineDocumento2 pagineNcm106 Course OutlineRS BuenavistaNessuna valutazione finora
- Jeong 2016Documento5 pagineJeong 2016francisco EscorciaNessuna valutazione finora
- VNG CB en Rev02 - HDDocumento5 pagineVNG CB en Rev02 - HDRudolfGerNessuna valutazione finora
- Prenatal DiagnosisDocumento11 paginePrenatal DiagnosisSonia BalintNessuna valutazione finora
- Ampathchat 45 Introducing The Iona NiptDocumento2 pagineAmpathchat 45 Introducing The Iona Niptalikiyaei82Nessuna valutazione finora
- NIV DEN KitDocumento1 paginaNIV DEN KitprudhviNessuna valutazione finora
- 03 Facchini - Hatch - India - Sep2023 - Shaping The Future - FinalDocumento33 pagine03 Facchini - Hatch - India - Sep2023 - Shaping The Future - FinalGlen ChoNessuna valutazione finora
- Science 01 April 2011Documento136 pagineScience 01 April 2011Mirsad Sommy Çoviç100% (1)
- POCKIT Central Nucleic Acid Analyzer (CE-IVD)Documento2 paginePOCKIT Central Nucleic Acid Analyzer (CE-IVD)Fadli AmbaraNessuna valutazione finora
- Brochure Prenatal TestingDocumento4 pagineBrochure Prenatal TestingMCuk2606Nessuna valutazione finora
- NeoBona BROCHURE PDFDocumento2 pagineNeoBona BROCHURE PDFyousrazeidan1979Nessuna valutazione finora
- EMVision Investor PresentationDocumento22 pagineEMVision Investor PresentationIlya HoffmanNessuna valutazione finora
- MoDiSe in M-Health SummitDocumento1 paginaMoDiSe in M-Health SummitAdesina IluyemiNessuna valutazione finora
- Thermo Scientific Refrigeradora PDFDocumento22 pagineThermo Scientific Refrigeradora PDFJulio David Vilca PizarroNessuna valutazione finora
- Cap - 03 Ciencia EndodonticaDocumento32 pagineCap - 03 Ciencia EndodonticaSuzana CunhaNessuna valutazione finora
- EZ1 Validation Guide: Sample & Assay TechnologiesDocumento28 pagineEZ1 Validation Guide: Sample & Assay TechnologiesJordy Daniel Vicente HernandezNessuna valutazione finora
- Ibs-108-G Dna Meth PN Final LowDocumento8 pagineIbs-108-G Dna Meth PN Final LowRohitNessuna valutazione finora
- New Surveys and Anatomic Pathology Education Programs For 2022 and 2023Documento2 pagineNew Surveys and Anatomic Pathology Education Programs For 2022 and 2023Amit BiswasNessuna valutazione finora
- Nanomaterials For Virus DetectionDocumento7 pagineNanomaterials For Virus DetectionRichard J. GrayNessuna valutazione finora
- Bombay Hospital InformationBookletDocumento12 pagineBombay Hospital InformationBookletmilkywayNessuna valutazione finora
- Hologic Discovery A BrochureDocumento8 pagineHologic Discovery A BrochureManuelNessuna valutazione finora
- Siemens Medical Solutions Molecular Imaging: Innovation Is in Our GenesDocumento18 pagineSiemens Medical Solutions Molecular Imaging: Innovation Is in Our GenesVĩ ĐặngNessuna valutazione finora
- 6 - Wilson Disease - ATI Active Learning Template System Disorder-1Documento1 pagina6 - Wilson Disease - ATI Active Learning Template System Disorder-1OlgaKNessuna valutazione finora
- Molecular Approaches To DiagnosingDocumento7 pagineMolecular Approaches To DiagnosingAkindele O AdigunNessuna valutazione finora
- Sage Trisomy Example Screening ReportDocumento2 pagineSage Trisomy Example Screening ReportpathbiomedxNessuna valutazione finora
- Mindray CLIA - SARS-CoV-2 IgM and IgG (June 2020)Documento7 pagineMindray CLIA - SARS-CoV-2 IgM and IgG (June 2020)enzaeniNessuna valutazione finora
- DR Gabriel Wallau Genomic Data Sharing Covid19 BrazilDocumento18 pagineDR Gabriel Wallau Genomic Data Sharing Covid19 Braziliq_dianaNessuna valutazione finora
- Xpert TVDocumento4 pagineXpert TVrhoderickNessuna valutazione finora
- Genomics in the AWS Cloud: Analyzing Genetic Code Using Amazon Web ServicesDa EverandGenomics in the AWS Cloud: Analyzing Genetic Code Using Amazon Web ServicesNessuna valutazione finora
- Binder 3 of 4 Dec-2018Documento1.169 pagineBinder 3 of 4 Dec-2018Anonymous OEmUQuNessuna valutazione finora
- Factors Affecting Physical FitnessDocumento7 pagineFactors Affecting Physical FitnessMary Joy Escanillas Gallardo100% (2)
- 40 RT-flex Control-System Rev01Documento68 pagine40 RT-flex Control-System Rev01Mayvon Botelho100% (2)
- SCIENCEEEEEDocumento3 pagineSCIENCEEEEEChristmae MaganteNessuna valutazione finora
- Rac Question PaperDocumento84 pagineRac Question PaperibrahimNessuna valutazione finora
- The Variable Resistor Has Been AdjustedDocumento3 pagineThe Variable Resistor Has Been AdjustedPank O RamaNessuna valutazione finora
- Chapter 1 (PLC)Documento9 pagineChapter 1 (PLC)Kibria PrangonNessuna valutazione finora
- Furuno CA 400Documento345 pagineFuruno CA 400Димон100% (3)
- Dialog Bahasa InggirsDocumento2 pagineDialog Bahasa InggirsKeRtha NeghaRaNessuna valutazione finora
- 09 Passport 7K 15K Performance Guidelines PCR 3 0Documento44 pagine09 Passport 7K 15K Performance Guidelines PCR 3 0thed719Nessuna valutazione finora
- Astm A194 2020Documento12 pagineAstm A194 2020rolando cuadro blancoNessuna valutazione finora
- S590 Machine SpecsDocumento6 pagineS590 Machine SpecsdilanNessuna valutazione finora
- Natural Disasters Vocabulary Exercises Fun Activities Games Icebreakers Oneonone Activiti 42747Documento2 pagineNatural Disasters Vocabulary Exercises Fun Activities Games Icebreakers Oneonone Activiti 42747Andrea Tercero VillarroelNessuna valutazione finora
- Sample Dilapidation ReportDocumento8 pagineSample Dilapidation ReportczarusNessuna valutazione finora
- Overall Method StatementDocumento33 pagineOverall Method Statementsaranga100% (1)
- Procter and Gamble-1Documento5 pagineProcter and Gamble-1Abegiel MendozaNessuna valutazione finora
- Chemistry Notes: SUBJECT: Leaving Cert Chemistry Level: TEACHER: Tara LyonsDocumento5 pagineChemistry Notes: SUBJECT: Leaving Cert Chemistry Level: TEACHER: Tara LyonsSevinc NuriyevaNessuna valutazione finora
- Smoldering Combustion: Guillermo ReinDocumento20 pagineSmoldering Combustion: Guillermo ReinAhmed HussainNessuna valutazione finora
- LighthouseDocumento4 pagineLighthousejaneborn5345Nessuna valutazione finora
- Donna Hay Magazine 2014-10-11 PDFDocumento172 pagineDonna Hay Magazine 2014-10-11 PDFlekovic_tanjaNessuna valutazione finora
- Viscous Fluid Flow Frank M White Third Edition - Compress PDFDocumento4 pagineViscous Fluid Flow Frank M White Third Edition - Compress PDFDenielNessuna valutazione finora
- Army Aviation Digest - Nov 1978Documento52 pagineArmy Aviation Digest - Nov 1978Aviation/Space History Library100% (1)
- 10th Aug. 2011 Structural Calculation (For Sub.) - 03Documento29 pagine10th Aug. 2011 Structural Calculation (For Sub.) - 03Nguyễn Tiến Việt100% (1)
- Case StudyDocumento61 pagineCase StudyA GNessuna valutazione finora
- Kuiz1 210114Documento12 pagineKuiz1 210114Vincent HoNessuna valutazione finora
- Biomediacal Waste Project FinalDocumento43 pagineBiomediacal Waste Project Finalashoknr100% (1)
- Tesla Magazine Vol4Documento48 pagineTesla Magazine Vol4jonathan100% (1)
- Course Structure and Content For Mechatronics, Systems and CDocumento32 pagineCourse Structure and Content For Mechatronics, Systems and CAnimonga HajimeNessuna valutazione finora
- LPPDocumento4 pagineLPPMargarida ReisNessuna valutazione finora
- MATLAB Fundamentals Quick ReferenceDocumento43 pagineMATLAB Fundamentals Quick ReferenceCarlos Manuel Cardoza EspitiaNessuna valutazione finora