Documenti di Didattica
Documenti di Professioni
Documenti di Cultura
Progress Dissecting The Oral Microbiome in Caries and Health
Caricato da
Jing XueTitolo originale
Copyright
Formati disponibili
Condividi questo documento
Condividi o incorpora il documento
Hai trovato utile questo documento?
Questo contenuto è inappropriato?
Segnala questo documentoCopyright:
Formati disponibili
Progress Dissecting The Oral Microbiome in Caries and Health
Caricato da
Jing XueCopyright:
Formati disponibili
Advanceshttp://adr.sagepub.
com/
in Dental Research
Progress Dissecting the Oral Microbiome in Caries and Health
R.A. Burne, L. Zeng, S.J. Ahn, S.R. Palmer, Y. Liu, T. Lefebure, M.J. Stanhope and M.M. Nascimento
ADR 2012 24: 77
DOI: 10.1177/0022034512449462
The online version of this article can be found at:
http://adr.sagepub.com/content/24/2/77
Published by:
http://www.sagepublications.com
On behalf of:
International and American Associations for Dental Research
Additional services and information for Advances in Dental Research can be found at:
Email Alerts: http://adr.sagepub.com/cgi/alerts
Subscriptions: http://adr.sagepub.com/subscriptions
Reprints: http://www.sagepub.com/journalsReprints.nav
Permissions: http://www.sagepub.com/journalsPermissions.nav
>> Version of Record - Aug 16, 2012
What is This?
Downloaded from adr.sagepub.com at Sichuan University on September 17, 2012 For personal use only. No other uses without permission.
2012 International & American Associations for Dental Research
Progress Dissecting the Oral Microbiome
in Caries and Health
to caries. Specifically, work by Marquis and others focused on the
R.A. Burne1,*, L. Zeng1, S.J. Ahn1, S.R. Palmer1, idea that S. mutans thrives in cariogenic plaque because it is better
Y. Liu1, T. Lefebure2, M.J. Stanhope2, able to tolerate, to grow, and to metabolize carbohydrate in a low
and M.M. Nascimento3 pH environment (Bender et al., 1986; Bender and Marquis, 1987;
Marquis, 1990; Belli and Marquis, 1991).
1
Department of Oral Biology, University of Florida College of Dentistry, Over the past decade or so, the application of DNA-based
1395 Center Drive, Gainesville, FL 32610, USA; 2Department of technologies to the analysis of the composition of the oral
Population Medicine and Diagnostic Sciences, Cornell University, Ithaca, microbiome in healthy individuals and in those with caries has
NY, USA; and 3Department of Restorative Dental Sciences, University of allowed for a more comprehensive cataloging of the species that
Florida College of Dentistry, Gainesville, FL, USA; *corresponding appear to be beneficial to dental health or to contribute to the
author, rburne@dental.ufl.edu
caries process, respectively.
Adv Dent Res 24(2):77-80, 2012 Oral health researchers have been world leaders in the appli-
cation of these types of technologies to the illumination of the
composition of the oral microbiome in health and diseasein
Abstract some cases, as a function of the hosts overall health. For a com-
Recent rapid advances in -omics technologies have yielded prehensive treatise on progress in the understanding of the oral
new insights into the interaction of the oral microbiome with its microbiome, readers are referred to a recent paper from the
host. Associations of species that are usually considered to be group at The Forsyth Institute (Dewhirst et al., 2010). In terms
acid-tolerant with caries have been confirmed, while some rec- of dental caries specifically, hybridization-based and sequence-
ognized as health-associated are often present in greater propor- based technologies have proven effective in developing a better
tions in the absence of caries. In addition, some newly identified picture of caries etiology, both from the standpoint of providing
bacteria have been suggested as potential contributors to the a more comprehensive understanding of the composition of the
caries process. In spite of this progress, two major challenges microbiome in caries and health, and by identifying species,
remain. The first is that there is a great deal of heterogeneity in some of them previously uncharacterized or uncultivated, as
the phenotypic capabilities of individual species of oral bacteria. possible contributors to the caries process. Some of these data
The second is that the most abundant taxa in oral biofilms dis- and more detail on the current view of caries and the oral micro-
play remarkable phenotypic plasticity, i.e., the bacteria associ- biome have been well-summarized in a recent review by How-
ated most strongly with health or with caries can morph rapidly ard Jenkinson (Jenkinson, 2011). In particular, the findings that
in response to alterations in environmental pH, carbohydrate S. mutans is not always isolable from active lesions and that the
availability and source, and oxygen tension and redox environ- proportions of certain species tend to be enhanced in health
ment. However, new technologic advances coupled with old- (e.g., S. sanguinis, S. gordonii), whereas caries lesions contain
fashioned microbiology are starting to erode the barriers to a higher proportions of aciduric S. mutans, non-mutans strepto-
more complete understanding of oral biofilm physiology and cocci, aciduric lactobacilli, bifidobacteria, Actinomyces spp.,
ecology, and in doing so are beginning to provide insights for and Scardovia spp., led Jenkinson to propose that pathogenic
the creation of novel cost-effective caries control therapies. communities rather than traditional microbial etiologies are
the drivers of dental caries. A general consensus seems to be
T he understanding of the etiology of dental caries in humans has
evolved and changed considerably over the last two decades.
It is now widely accepted that dental caries occurs when envi-
developing that this is a reasonable way to view the microbiol-
ogy of dental caries.
ronmental pressures, primarily in the form of frequent intake of
dietary carbohydrate and repeated acidification of oral biofilms,
The Known Unknowns
lead to changes in the proportions of particular bacterial species as The application of 16S rRNA gene sequencing to analysis of the
a caries lesion is initiated and progresses. Arguably, a key concep- oral microbiome will continue to contribute to our knowledge of
tual advance that led to the current ecological view of dental caries
development was the recognition of the importance of the prop-
erty of aciduricity (acid tolerance) in S. mutans and its relationship Key Words
DOI: 10.1177/0022034512449462 biofilm, 16S rRNA, acid tolerance, arginine metabolism, ecology,
pH homeostasis.
International & American Associations for Dental Research
77
Downloaded from adr.sagepub.com at Sichuan University on September 17, 2012 For personal use only. No other uses without permission.
2012 International & American Associations for Dental Research
78 Burne et al. Adv Dent Res 24(2) 2012
Table. Overview of Phenotypic Properties of 16 S. mutans Isolates for today is to correlate genotypic data effectively with the pheno-
Which a High-coverage DNA Sequence Is Now Available typic capacities of organisms and clinical status of the individu-
Condition Doubling Time (Range in min) als. Said another way, microbiologists need to develop the
knowledge base to make reliable correlations of the gene con-
Normal1 65-125
tent of individual species with the ability of oral commensals to
pH 5.52 164-328
enhance the stability and beneficial nature of oral biofilms or the
Aerated3 87-218
Paraquat4 105-387 (1 NG) 5 capacity of cariogenic organisms to damage host tissues.
1
Growth in static culture in Brain Heart Infusion broth in a 5% CO2
atmosphere. Exploring the Relationship of
2
Growth in static culture in a 5% CO2 atmosphere in Brain Heart
Infusion broth that was acidified to pH 5.5. Genotype and Cariogenic Potential in
3
Growth in Brain Heart Infusion broth with shaking at 150 RPM in an Streptococcus mutans
aerobic environment.
4
Growth in Brain Heart Infusion broth in static culture with 25 mM Numerous studies have demonstrated substantial genetic heteroge-
paraquat. neity in clinical isolates of S. mutans. [For recent examples, see
5
NG = No growth. One isolate was unable to initiate any growth in
the presence of paraquat. Zhang et al. (2009), Arthur et al. (2011), Cheon et al. (2011), and
Phattarataratip et al. (2011).] Although these studies have enhanced
our basic understanding of the biology and epidemiology of
the oral microbiome, and with new technologies, the depth and S. mutans, our present knowledge of gene content across the spe-
breadth of the information collected should be remarkable. But cies and its relationship to phenotypes is minimal. As part of a
this technology is not a panacea, and there is a very good chance collaborative project between the University of Florida and Cornell
that such studies will prove confirmatory at best. In fact, it may University, we have completed a high-coverage DNA sequence of
be that such studies serve to further complicate the interpretation 57 genomes of S. mutans, as well as genomes of the closely related
of the relationships of different bacteria with caries and health. mutans streptococci S. ratti and S. cricetus. A manuscript detailing
For example, in a study by Griffen and co-workers, species that the gene content and evolutionary biology has been submitted, and
were previously thought to be associated primarily with health all sequences will be posted to a convenient Web browser. One of
were also elevated in caries-active individuals, leading the the goals of this undertaking was to begin to correlate gene content
authors to conclude that The relationship of acid-base metabo- with phenotype, which we have undertaken by initiating pheno-
lism to 16S rRNA gene-based species assignments appears to be typic characterization of 16 genetically diverse isolates of S.
complex (Gross et al., 2010). This conclusion confirms mutans for which we have the high-coverage sequence. The Table
what many microbiologists who study oral bacteria have known shows the results of analysis of a selected group of phenotypic
for a long time: Individual species of oral commensals and properties that highlights the diversity of this species. Collectively,
pathogens display a remarkable degree of genetic heterogeneity. the genetic diversity observed from analysis of a whole-genome
The ramifications of this genetic heterogeneity in the context of sequence of these strains of S. mutans is mirrored by a high degree
caries etiology are that one cannot rely solely on 16S sequence of phenotypic diversity. Importantly, there is diversity in traits that
information to understand the cariogenic potential of the oral directly affect the establishment, persistence, and cariogenic
microbiome. This is because individual isolates of S. mutans, potential of S. mutans in humans. Studies are ongoing to explore
oral commensal streptococci, and other common dental plaque correlations of gene content with critical phenotypic traits.
bacteria differ widely in those characteristicsadherence, bio-
film formation, acidogenesis, and acid tolerancethat have a Understanding Ph Homeostasis in Health-
direct impact on their ability to contribute to the caries process.
Another reason why the 16S sequence alone appears inade-
Associated Dental Biofilms
quate to predict associations of particular bacteria with oral Embodied in the ecological plaque hypothesis (Marsh, 2009)
diseases is related to the concept of phenotypic plasticity. That for dental caries is the concept that certain functions are well-
is, depending on the environment in which the organisms are represented in health-associated biofilms, whereas certain other
growing, the phenotypes of the bacteria can be quite different functions, such as acidogenesis and acid tolerance, are better
(Lemos and Burne, 2008). Environmental factors, including pH, represented in biofilms on or near active caries lesions. Since
oxygen and redox, carbohydrate source and availability, popula- caries is a disease that does not occur without the production of
tion density, and the growth phase of the bacteria, have been a low pH environment by oral biofilms, one might predict that
shown to have a profound impact on the gene expression pro- activities that help to promote a more neutral pH could have a
files of many oral streptococci and to influence phenotypes that strong inhibitory effect on caries development. It is known that
directly influence cariogenicity (Burne et al., 2009). Therefore, there are two substrates provided in saliva and the diet that are
the cariogenic potential of particular strains of commensals or particularly effective at eliciting an increase in the pH when
pathogens could be greatly enhanced in vivo given the right host metabolized by populations of oral bacteria: urea and arginine
environment, making it impossible to predict solely from 16S (Fig.). A more comprehensive review of oral arginine and urea
sequence or using simple in vitro assays which bacteria have the metabolism is provided elsewhere (Burne and Marquis, 2000).
greatest potential to damage dental tissue. In light of these findings, Here we focus on new information related to arginine metabo-
one can posit that the most crucial goal for caries microbiology lism and caries resistance in humans.
Downloaded from adr.sagepub.com at Sichuan University on September 17, 2012 For personal use only. No other uses without permission.
2012 International & American Associations for Dental Research
Adv Dent Res 24(2) 2012 Genetic and Phenotypic Heterogeneity in Oral Biofilms 79
Arginine is metabolized by oral biofilms primarily by a
3-enzyme pathway called the arginine deiminase system (ADS).
The net reaction is that arginine is converted to ornithine and 2
molecules of ammonia, which causes the pH to rise. The bacte-
ria also gain an ATP in the process (Burne and Marquis, 2000).
Thus, metabolism of arginine via the ADS is beneficial to the
host because it raises biofilm pH, which discourages the out-
growth of cariogenic species and reduces damage to enamel.
The ADS is also beneficial to the bacteria that have it because
the ammonia it generates neutralizes the cytoplasm of the bacte-
ria, which protects them from acid damage, and the ATP pro-
duced can be used for growth or homeostasis.
One major impetus for generating a better understanding of the Figure. The principle sources of, and pathways for, pH homeostasis in
ADS in oral bacteria was the idea that enhancing ADS activity in oral biofilms. Urea is present in all salivary gland secretions and in gin-
dental plaque could have a beneficial effect on biofilm ecology gival crevicular fluid at concentrations approximating those in serum
and provide protection against caries. It is only recently, though, (1-10 mM). Arginine is present in low M quantities in salivary secre-
tions, but is abundant in peptide form. Urease enzymes are encoded
that the arginolytic capacity of the bacteria in dental plaque and
by a relatively small number of oral bacteria, mainly Streptococcus
saliva has been compared in caries-active and caries-free indi- salivarius, some Actinomyces spp., and oral Haemophilus spp., and
viduals. Consistent with its predicted role, individuals who had no cleave urea into 2 molecules of ammonia and 1 of CO2. The arginine
clinical evidence of past or present caries activity had signifi- deiminase pathway (ADS) is a 3-enzyme system that cleaves arginine
cantly higher levels of the arginine deiminase enzyme measurable to ornithine, 2 molecules of ammonia, and 1 of CO2. The most abun-
in their plaque and salivary bacterial populations compared with dant ADS-positive bacteria are oral Streptococcus spp. and some lacto-
those who had active caries lesions (Nascimento et al., 2009). bacilli, including Lactobacillus fermentum. The oral streptococci tend to
express the ADS at much higher levels than other oral bacteria.
This seminal study was conducted with plaque that had been
pooled from different sites in adult humans, but more recently,
Nascimento and co-workers have shown that ADS activity in
plaques isolated from carious surfaces in children is markedly
lower than that from sound surfaces. Notably, the range of ADS Recently, we have begun to investigate the molecular basis
activity among these children was greater than 10,000-fold, so for variability in the expression of the ADS in these isolates.
there is huge variation in the arginolytic potential of plaque in What is emerging is that the differences are, in part, associated
children. Given that we have shown in vitro that the ability of with the absolute capacity of the bacteria to produce the
ammonia production by bacteria to temper acidification during a enzymes, but are also strongly related to the way the ADS genes
glucose challenge is directly correlated with the quantity of ADS are regulated in response to environmental factors. Extrapolat-
enzymes present in the bacteria, these findings have tremendous ing these findings to the in vivo setting, one may conclude that
implications for caries control. However, more information on the there are certain isolates of common commensal strains of oral
organisms responsible for arginolysis in dental plaque is needed. streptococci that have a high constitutional capacity to provide
It is here that the heterogeneity of human oral bacteria protection against caries. Rapid methods to discriminate highly
becomes particularly germane and interesting. Specifically, arginolytic isolates from weakly ADS-positive strains could be
Nascimento and co-workers, using PCR-based methods, a very powerful addition to caries risk assessment protocols.
explored whether dental plaque that had low arginolytic capac-
ity also had decreased proportions of the most common ADS-
Summary
positive bacteria (Nascimento et al., 2009). Surprisingly, high
ADS activity could not be correlated with high numbers of the Technological advances in the use of high-throughput sequenc-
more prominent known ADS-positive bacteria. After additional ing platforms for analysis of the composition of the oral micro-
experimentation, it was determined that there is remarkable biome, coupled with traditional microbiological studies of the
heterogeneity within the species of bacteria that possess the organisms isolated from human dental plaque, have provided a
ADS, vis--vis their ability to produce high levels of the new framework for interpreting caries etiology. However,
enzymes. Interestingly, within a given species, a wide range of genetic diversity and complexities in the adaptive strategies of
abilities to break down arginine was noted (manuscript in prepa- oral bacteria to fluctuating biofilm conditions diminish the util-
ration). For example, individual isolates of S. sanguinis differed ity of taxa-level identification of the microbiome for developing
by as much as 100-fold in the levels of AD enzyme activity. correlations of populations of specific organisms with health,
Therefore, intraspecies variability in the capacity to break down especially, but also with caries. The marriage of functional stud-
arginine may account, in large part, for the variability in ADS ies of the microbiome with whole-genome and phenotypic
activity in caries-active and -free individuals, reinforcing the analyses of multiple species and multiple isolates from individu-
idea that species- or taxa-level identification of organisms in the als of known caries status is needed to yield the level of knowl-
microbiome is not adequate to come to terms with the patho- edge that will facilitate the rational design of preventive and
genic potential of oral biofilms. treatment strategies to eradicate dental caries on a global scale.
Downloaded from adr.sagepub.com at Sichuan University on September 17, 2012 For personal use only. No other uses without permission.
2012 International & American Associations for Dental Research
80 Burne et al. Adv Dent Res 24(2) 2012
Acknowledgments Cheon K, Moser SA, Whiddon J, Osgood RC, Momeni S, Ruby JD, et al.
(2011). Genetic diversity of plaque mutans streptococci with rep-PCR.
This work was supported by the NIH-NIDCR (DE10362), the J Dent Res 90:331-335.
NIH-NIAID (AI073368), and the Colgate-Palmolive Co. The Dewhirst FE, Chen T, Izard J, Paster BJ, Tanner AC, Yu WH, et al. (2010).
The human oral microbiome. J Bacteriol 192:5002-5017.
authors declare no potential conflicts of interest with respect to Gross EL, Leys EJ, Gasparovich SR, Firestone ND, Schwartzbaum JA,
the authorship and/or publication of this article. Janies DA, et al. (2010). Bacterial 16S sequence analysis of severe car-
ies in young permanent teeth. J Clin Microbiol 48:4121-4128.
Jenkinson HF (2011). Beyond the oral microbiome. Environ Microbiol
References 13:3077-3087.
Lemos JA, Burne RA (2008). A model of efficiency: stress tolerance by
Arthur RA, Cury AA, Graner RO, Rosalen PL, Vale GC, Paes Leme AF, Streptococcus mutans. Microbiology 154(Pt 11):3247-3255.
et al. (2011). Genotypic and phenotypic analysis of S. mutans isolated Marquis RE (1990). Diminished acid tolerance of plaque bacteria caused by
from dental biofilms formed in vivo under high cariogenic conditions. fluoride. J Dent Res 69(Spec Iss):672-675.
Braz Dent J 22:267-274. Marsh PD (2009). Dental plaque as a biofilm: the significance of pH in
Belli WA, Marquis RE (1991). Adaptation of Streptococcus mutans and health and caries. Compend Contin Educ Dent 30:76-87.
Enterococcus hirae to acid stress in continuous culture. Appl Environ Nascimento MM, Gordan VV, Garvan CW, Browngardt CM, Burne RA
Microbiol 57:1134-1138. (2009). Correlations of oral bacterial arginine and urea catabolism with
Bender GR, Marquis RE (1987). Membrane ATPases and acid tolerance of caries experience. Oral Microbiol Immunol 24:89-95.
Actinomyces viscosus and Lactobacillus casei. Appl Environ Microbiol Phattarataratip E, Olson B, Broffitt B, Qian F, Brogden KA, Drake DR,
53:2124-2128. et al. (2011). Streptococcus mutans strains recovered from caries-active
Bender GR, Sutton SV, Marquis RE (1986). Acid tolerance, proton permeabil- or caries-free individuals differ in sensitivity to host antimicrobial pep-
ities, membrane ATPases of oral streptococci. Infect Immun 53:331-338. tides. Mol Oral Microbiol 26:187-199.
Burne RA, Marquis RE (2000). Alkali production by oral bacteria and pro- Zhang L, Foxman B, Drake DR, Srinivasan U, Henderson J, Olson B, et al.
tection against dental caries. FEMS Microbiol Lett 193:1-6. (2009). Comparative whole-genome analysis of Streptococcus mutans
Burne RA, Ahn SJ, Wen ZT, Zeng L, Lemos JA, Abranches J, et al. (2009). isolates within and among individuals of different caries status. Oral
Opportunities for disrupting cariogenic biofilms. Adv Dent Res 21:17-20. Microbiol Immunol 24:197-203.
Downloaded from adr.sagepub.com at Sichuan University on September 17, 2012 For personal use only. No other uses without permission.
2012 International & American Associations for Dental Research
Potrebbero piacerti anche
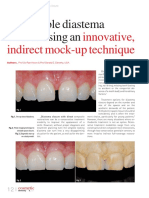
- Innovative, Indirect Mock-Up Technique: Predictable Diastema Closure Using AnDocumento4 pagineInnovative, Indirect Mock-Up Technique: Predictable Diastema Closure Using AnJing XueNessuna valutazione finora
- 10.2319 080619-516EPubDocumento8 pagine10.2319 080619-516EPubJing XueNessuna valutazione finora
- Case Report: Diastema Closure in Anterior Teeth Using A Posterior MatrixDocumento7 pagineCase Report: Diastema Closure in Anterior Teeth Using A Posterior MatrixJing XueNessuna valutazione finora
- Pak Orthod J 2013 5 1 27 33 PDFDocumento7 paginePak Orthod J 2013 5 1 27 33 PDFrief_candidaNessuna valutazione finora
- ContentServer PDFDocumento7 pagineContentServer PDFArchidHananAfifahNessuna valutazione finora
- 10 1111@jopr 13109Documento6 pagine10 1111@jopr 13109Jing XueNessuna valutazione finora
- 33-Article Text-71-1-10-20120907Documento5 pagine33-Article Text-71-1-10-20120907Jing XueNessuna valutazione finora
- 10-Pramod Philip (Short Communication)Documento3 pagine10-Pramod Philip (Short Communication)Jing XueNessuna valutazione finora
- 5-BarrosdeCamposetal 2015IJEDDocumento12 pagine5-BarrosdeCamposetal 2015IJEDJing XueNessuna valutazione finora
- Closure of Diastema and Gingival Recontouring Using Direct Adhesive Restorations: A Case ReportDocumento12 pagineClosure of Diastema and Gingival Recontouring Using Direct Adhesive Restorations: A Case ReportJing XueNessuna valutazione finora
- Conservative Approach For The Esthetic Management of Multiple Interdental Spaces: A Systematic ApproachDocumento11 pagineConservative Approach For The Esthetic Management of Multiple Interdental Spaces: A Systematic ApproachJing XueNessuna valutazione finora
- 10 1111@jopr 13109Documento6 pagine10 1111@jopr 13109Jing XueNessuna valutazione finora
- Midline Diastema Closure With Partial Laminate Veneers: A Case ReportDocumento4 pagineMidline Diastema Closure With Partial Laminate Veneers: A Case ReportAndykaYayanSetiawanNessuna valutazione finora
- 0003-3219 (1977) 047 0265 Tmidai 2 0 Co 2Documento7 pagine0003-3219 (1977) 047 0265 Tmidai 2 0 Co 2Jing XueNessuna valutazione finora
- Factors Affecting HealingDocumento12 pagineFactors Affecting HealingpriyadharshiniNessuna valutazione finora
- Survival of Restored Endodontically Treated Teeth in Relation To Periodontal StatusDocumento4 pagineSurvival of Restored Endodontically Treated Teeth in Relation To Periodontal StatusJing XueNessuna valutazione finora
- Intraorifice Sealing Ability of Different Materials in Endodontically Treated TeethDocumento5 pagineIntraorifice Sealing Ability of Different Materials in Endodontically Treated TeethJing XueNessuna valutazione finora
- Midline Diastema Closure With Partial Laminate Veneers: A Case ReportDocumento4 pagineMidline Diastema Closure With Partial Laminate Veneers: A Case ReportAndykaYayanSetiawanNessuna valutazione finora
- Incidence of Pain Associated With Clinical Factors During and After Root Canal Therapy. Part 1. Interappointment PainDocumento4 pagineIncidence of Pain Associated With Clinical Factors During and After Root Canal Therapy. Part 1. Interappointment PainJing XueNessuna valutazione finora
- Do We Really Know How To Evaluate Tooth Prognosis A Systematic Review and Suggested Approach.1 PDFDocumento11 pagineDo We Really Know How To Evaluate Tooth Prognosis A Systematic Review and Suggested Approach.1 PDFJing XueNessuna valutazione finora
- Wide Diameter Implants-Degidi Piattelli LezziDocumento7 pagineWide Diameter Implants-Degidi Piattelli LezziJing XueNessuna valutazione finora
- Outcome of Root Canal Treatment in Dogs Determined by Periapical Radiography and Cone-Beam Computed Tomography ScansDocumento4 pagineOutcome of Root Canal Treatment in Dogs Determined by Periapical Radiography and Cone-Beam Computed Tomography ScansJing XueNessuna valutazione finora
- Alve Olar Changes in Sockets Post ExtractionDocumento11 pagineAlve Olar Changes in Sockets Post ExtractionPratyusha VallamNessuna valutazione finora
- Dental Vertical Root Fractures: Value of CT in Detection: Head and Neck ImagingDocumento5 pagineDental Vertical Root Fractures: Value of CT in Detection: Head and Neck ImagingJing XueNessuna valutazione finora
- When To Refer To Specialist3Documento2 pagineWhen To Refer To Specialist3Jing XueNessuna valutazione finora
- Walton SinusTractTracingEffectivenessDocumento1 paginaWalton SinusTractTracingEffectivenessJing XueNessuna valutazione finora
- Why Clinicians Are Natural Bayesians: Interpreting Diagnostic Test Results From The Bayesian PerspectiveDocumento4 pagineWhy Clinicians Are Natural Bayesians: Interpreting Diagnostic Test Results From The Bayesian PerspectiveJing XueNessuna valutazione finora
- When To Refer To Specialist2Documento2 pagineWhen To Refer To Specialist2Jing XueNessuna valutazione finora
- Project Deliverable: Safety and Efficacy of A New and Emerging Dental X-Ray ModalityDocumento16 pagineProject Deliverable: Safety and Efficacy of A New and Emerging Dental X-Ray ModalityJing XueNessuna valutazione finora
- Avulsions and Intrusions: The Controversial Displacement InjuriesDocumento7 pagineAvulsions and Intrusions: The Controversial Displacement InjuriesJing XueNessuna valutazione finora
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeDa EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeValutazione: 4 su 5 stelle4/5 (5794)
- The Little Book of Hygge: Danish Secrets to Happy LivingDa EverandThe Little Book of Hygge: Danish Secrets to Happy LivingValutazione: 3.5 su 5 stelle3.5/5 (400)
- Shoe Dog: A Memoir by the Creator of NikeDa EverandShoe Dog: A Memoir by the Creator of NikeValutazione: 4.5 su 5 stelle4.5/5 (537)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceDa EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceValutazione: 4 su 5 stelle4/5 (895)
- The Yellow House: A Memoir (2019 National Book Award Winner)Da EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Valutazione: 4 su 5 stelle4/5 (98)
- The Emperor of All Maladies: A Biography of CancerDa EverandThe Emperor of All Maladies: A Biography of CancerValutazione: 4.5 su 5 stelle4.5/5 (271)
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryDa EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryValutazione: 3.5 su 5 stelle3.5/5 (231)
- Never Split the Difference: Negotiating As If Your Life Depended On ItDa EverandNever Split the Difference: Negotiating As If Your Life Depended On ItValutazione: 4.5 su 5 stelle4.5/5 (838)
- Grit: The Power of Passion and PerseveranceDa EverandGrit: The Power of Passion and PerseveranceValutazione: 4 su 5 stelle4/5 (588)
- On Fire: The (Burning) Case for a Green New DealDa EverandOn Fire: The (Burning) Case for a Green New DealValutazione: 4 su 5 stelle4/5 (74)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureDa EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureValutazione: 4.5 su 5 stelle4.5/5 (474)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaDa EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaValutazione: 4.5 su 5 stelle4.5/5 (266)
- The Unwinding: An Inner History of the New AmericaDa EverandThe Unwinding: An Inner History of the New AmericaValutazione: 4 su 5 stelle4/5 (45)
- Team of Rivals: The Political Genius of Abraham LincolnDa EverandTeam of Rivals: The Political Genius of Abraham LincolnValutazione: 4.5 su 5 stelle4.5/5 (234)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyDa EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyValutazione: 3.5 su 5 stelle3.5/5 (2259)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreDa EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreValutazione: 4 su 5 stelle4/5 (1090)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersDa EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersValutazione: 4.5 su 5 stelle4.5/5 (344)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)Da EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Valutazione: 4.5 su 5 stelle4.5/5 (121)
- Her Body and Other Parties: StoriesDa EverandHer Body and Other Parties: StoriesValutazione: 4 su 5 stelle4/5 (821)
- A Detailed Lesson Plan - The Fundamental Law of ProportionDocumento10 pagineA Detailed Lesson Plan - The Fundamental Law of ProportionPrincess De LeonNessuna valutazione finora
- MCS 150 Page 1Documento2 pagineMCS 150 Page 1Galina NovahovaNessuna valutazione finora
- Muscle Building MythsDocumento5 pagineMuscle Building MythsKarolNessuna valutazione finora
- Computer Awareness: Special Edition E-BookDocumento54 pagineComputer Awareness: Special Edition E-BookTanujit SahaNessuna valutazione finora
- LPP-Graphical and Simplex MethodDocumento23 pagineLPP-Graphical and Simplex MethodTushar DhandeNessuna valutazione finora
- Upload A Document To Access Your Download: Social Studies of HealthDocumento3 pagineUpload A Document To Access Your Download: Social Studies of Health1filicupENessuna valutazione finora
- Medical Secretary: A. Duties and TasksDocumento3 pagineMedical Secretary: A. Duties and TasksNoona PlaysNessuna valutazione finora
- On Evil - Terry EagletonDocumento44 pagineOn Evil - Terry EagletonconelcaballocansadoNessuna valutazione finora
- LAID BACK - Sunshine ReggaeDocumento16 pagineLAID BACK - Sunshine ReggaePablo ValdezNessuna valutazione finora
- The Nervous System 1ae60 62e99ab3Documento1 paginaThe Nervous System 1ae60 62e99ab3shamshadNessuna valutazione finora
- The Old Rugged Cross - George Bennard: RefrainDocumento5 pagineThe Old Rugged Cross - George Bennard: RefrainwilsonNessuna valutazione finora
- Gardens Illustrated FernandoDocumento4 pagineGardens Illustrated FernandoMariaNessuna valutazione finora
- Gmail - Payment Received From Cnautotool - Com (Order No - Cnautot2020062813795)Documento2 pagineGmail - Payment Received From Cnautotool - Com (Order No - Cnautot2020062813795)Luis Gustavo Escobar MachadoNessuna valutazione finora
- Unit 2 Lab Manual ChemistryDocumento9 pagineUnit 2 Lab Manual ChemistryAldayne ParkesNessuna valutazione finora
- Injection Pump Test SpecificationsDocumento3 pagineInjection Pump Test Specificationsadmin tigasaudaraNessuna valutazione finora
- Palladium Skill Book 1a.Documento10 paginePalladium Skill Book 1a.Randy DavisNessuna valutazione finora
- Fini Cat K-Max 45-90 enDocumento16 pagineFini Cat K-Max 45-90 enbujin.gym.essenNessuna valutazione finora
- Philosophies PrinceDocumento4 paginePhilosophies PrincePrince CuetoNessuna valutazione finora
- Catalogue of Khalsa Darbar Records Vol.1 - Compiled by Sita Ram KohliDocumento180 pagineCatalogue of Khalsa Darbar Records Vol.1 - Compiled by Sita Ram KohliSikhDigitalLibrary100% (1)
- HDFCDocumento60 pagineHDFCPukhraj GehlotNessuna valutazione finora
- Factoring Problems-SolutionsDocumento11 pagineFactoring Problems-SolutionsChinmayee ChoudhuryNessuna valutazione finora
- The Library PK - Library of Urdu BooksDocumento8 pagineThe Library PK - Library of Urdu Bookszamin4pakNessuna valutazione finora
- SMA Releases Online Version of Sunny Design For PV System ConfigurationDocumento3 pagineSMA Releases Online Version of Sunny Design For PV System ConfigurationlakliraNessuna valutazione finora
- Third Party Intervention in The Criminal TrialDocumento8 pagineThird Party Intervention in The Criminal TrialVenkat Raman JNessuna valutazione finora
- How To Become A Skilled OratorDocumento5 pagineHow To Become A Skilled OratorDonain Alexis CamarenaNessuna valutazione finora
- Fundamentals of Group DynamicsDocumento12 pagineFundamentals of Group DynamicsLimbasam PapaNessuna valutazione finora
- Hop Movie WorksheetDocumento3 pagineHop Movie WorksheetMARIA RIERA PRATSNessuna valutazione finora
- Shaping School Culture Case StudyDocumento7 pagineShaping School Culture Case Studyapi-524477308Nessuna valutazione finora
- CBLM FBS NC-II (Develop and Update Food and Beverage Service)Documento24 pagineCBLM FBS NC-II (Develop and Update Food and Beverage Service)Angel PanganibanNessuna valutazione finora
- Full Download Test Bank For Amgov Long Story Short 1st Edition Christine Barbour PDF Full ChapterDocumento13 pagineFull Download Test Bank For Amgov Long Story Short 1st Edition Christine Barbour PDF Full Chaptertithly.decamplh56c7100% (20)