Documenti di Didattica
Documenti di Professioni
Documenti di Cultura
Recent Trends in in Vitro Nanodiagnostics For Detection of PathogensReview Article
Caricato da
yyuipTitolo originale
Copyright
Formati disponibili
Condividi questo documento
Condividi o incorpora il documento
Hai trovato utile questo documento?
Questo contenuto è inappropriato?
Segnala questo documentoCopyright:
Formati disponibili
Recent Trends in in Vitro Nanodiagnostics For Detection of PathogensReview Article
Caricato da
yyuipCopyright:
Formati disponibili
Journal of Controlled Release 159 (2012) 164180
Contents lists available at SciVerse ScienceDirect
Journal of Controlled Release
journal homepage: www.elsevier.com/locate/jconrel
Review
Recent trends in in-vitro nanodiagnostics for detection of pathogens
Siddhesh B. Shinde, Clara B. Fernandes, Vandana B. Patravale
Institute of Chemical Technology, Matunga, Mumbai 400019, India
a r t i c l e
i n f o
Article history:
Received 21 June 2011
Accepted 23 November 2011
Available online 13 December 2011
Keywords:
Nanodiagnostics
Bacterial
Dendrimer
Quantum dots
Liposomes
Virus
a b s t r a c t
In recent times, infectious diseases are posing to be a major healthcare issue. The contagious nature of these
diseases makes it imperative to develop logical solutions for early diagnosis and containment of these diseases.
Traditional techniques suffer from limitations, including laborious sample preparation, bulky instrumentation
and slow data readout. In view of the urgency for sensitive, specic, robust and rapid diagnostics, numerous advancements have been made in the area of diagnostics. Broadly, most of these innovative approaches, have utilized the unique properties of nanomaterials in order to achieve detection of infectious agents, even in complex
media like blood, urine. Additionally, researchers have also leveraged the advantages of nanomaterials by coupling with novel methods, such as surface plasmon spectroscopy, amperometry and magnetic relaxation. Thus,
this review intends to provide an overview on nanotechnology-based in-vitro diagnostics encompassing the useful role of both bioreceptors and transducers in the diagnosis of infectious diseases in diverse settings throughout
the globe, preventing epidemics and safeguarding human and economic wellness.
2011 Elsevier B.V. All rights reserved.
Contents
1.
2.
3.
4.
General introduction . . . . . . . . . . . . . . . . .
1.1.
Traditional diagnostic techniques . . . . . . . .
1.1.1.
Isolation, growth and microscopy . . .
1.1.2.
Immunology-based methods . . . . . .
1.1.3.
Polymerase chain reaction (PCR) . . . .
1.1.4.
Miscellaneous techniques . . . . . . .
Potential of nanotechnology . . . . . . . . . . . . .
Nanotechnology based bioreceptors . . . . . . . . . .
3.1.
Liposomes . . . . . . . . . . . . . . . . . .
3.2.
Carbon nanotubes . . . . . . . . . . . . . . .
3.3.
Dendrimers . . . . . . . . . . . . . . . . . .
3.4.
Gold nanoparticles (AuNPs) . . . . . . . . . .
3.5.
Conducting polymeric nanoparticles . . . . . .
3.6.
Polystyrene nanoparticles . . . . . . . . . . .
3.7.
Iron oxide nanoparticles . . . . . . . . . . . .
3.8.
Quantum dots (QDs) . . . . . . . . . . . . . .
3.9.
Bacteriophage/virus particles . . . . . . . . . .
Nanotechnology based transducers . . . . . . . . . .
4.1.
Optical biosensors . . . . . . . . . . . . . . .
4.1.1.
Fluorescence-based detection . . . . .
4.1.2.
Refractive index detection . . . . . . .
4.2.
Electrochemical biosensors . . . . . . . . . . .
4.2.1.
Anodic stripping voltammetry (ASV) . .
4.2.2.
Amperometric sensors . . . . . . . .
4.2.3.
Potentiometric sensors . . . . . . . .
4.2.4.
Conductometric sensors . . . . . . . .
4.2.5.
Electrochemical impedance spectroscopy
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
. . .
(EIS)
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
165
165
165
165
165
166
166
166
166
167
167
168
168
168
169
169
169
170
170
170
171
171
172
172
172
173
173
Corresponding author at: Department of Pharmaceutical Sciences and Technology, Institute of Chemical Technology, Matunga, Mumbai-400019, India. Tel.: + 91 22 3361 2217;
fax: + 91 22 3361 1020.
E-mail address: vbp_muict@yahoo.co.in (V.B. Patravale).
0168-3659/$ see front matter 2011 Elsevier B.V. All rights reserved.
doi:10.1016/j.jconrel.2011.11.033
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
4.3.
Microgravimetric (Piezoelectrical) . . . . . . . . . . . . . . . .
4.3.1.
Bulk wave (BW) or Quartz crystal microbalance (QCM) . .
4.3.2.
Surface acoustic wave (SAW) devices . . . . . . . . . .
4.3.3.
Magnetoelastic sensors . . . . . . . . . . . . . . . . .
4.4.
Magnetic biosensors . . . . . . . . . . . . . . . . . . . . . .
4.4.1.
NMR technique . . . . . . . . . . . . . . . . . . . .
4.4.2.
Magnetic eld sensors based on superconducting quantum
4.4.3.
Magnetoresistive-based biosensors . . . . . . . . . . .
5.
Future prospects . . . . . . . . . . . . . . . . . . . . . . . . . . . .
Acknowledgement . . . . . . . . . . . . . . . . . . . . . . . . . . . . .
References . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . . .
1. General introduction
In today's changing scenarios, Prevention Is Better Than Cure the
paradigm has increasingly become hard-hitting truth of life [1,2]. The
capricious political situations have led to an increase in threat of potential biological weapons endangering the survival of mankind.
Broadly, these bio-chemical agents, Bacillus anthracis (anthrax), Variola major virus (small pox), Botulinum neurotoxins, Yersinia pestis,
Francisella tularensis, Burkholderia pseudomellei, Burkholderia mallei,
Rickettsia spp, Coxiella burnetti, Venezuelan equine encephalitis virus,
Marburg and Ebola viruses and inuenza viruses are considered to
have the great potential for mass causalities and civil disruption
[3,4]. Besides these, there are several emerging infectious diseases
with the potential for signicant public health consequences, including
malaria, HIV virus related complications, Dengue fever, West Nile fever,
and Rift Valley fever as well as tuberculosis [4]. As with biowarfare
agents, emerging infectious disease agents may be directly transmissible or vector borne. Further, infectious diseases prevalent in the developing world result in increased rates of morbidity and mortality.
While infectious diseases can initiate in a localized region, they can
spread rapidly at any moment due to the ease of traveling from one
part of the world to the next.
Because of the threat posed by both biowarfare agents and emerging or reemerging infectious disease agents, there is a need to develop
diagnostics for rapid identication of such agents in clinical setting in
order to treat the individuals at risk and to improve public health surveillance and epidemiology.
1.1. Traditional diagnostic techniques
1.1.1. Isolation, growth and microscopy
Traditionally, the presence of most pathogens such as bacteria,
fungi, protozoa and worms is determined microscopically, usually
after growth in a pure culture. These methods, although highly specific, have several limitations, to name a few; it is only useful for the
analysis of the samples having high pathogen load. It is a time consuming procedure (23 days for initial results, and up to 710 days
for conrmation) as growth pattern methods usually require the
growth of the pathogen in a particular medium followed by at least
a 24-h incubation period to yield results. It could result in false negative results especially in cases where some viable bacterial strains in
the environment can enter a dormancy state rendering them nonculturable (viable-but non-culturable (VBNC)) subsequently leading to
an underestimation of pathogen numbers or a failure to isolate and
identify a pathogen from a contaminated sample. These limitations
are even more signicant in the identication of viruses that due to
their small size (approx. 100 nm) cannot be studied using conventional optical microscopy and require the use of an electron microscope for their visualization. Moreover, prior to analysis, the culture
and growth of viruses in the laboratory necessitate use of extensive
protocols for their growth. Inspite of their disadvantages, conventional culture methods still represent a eld where progress is possible.
These methods are often combined together with other pathogen
. . . . . .
. . . . . .
. . . . . .
. . . . . .
. . . . . .
. . . . . .
interference
. . . . . .
. . . . . .
. . . . . .
. . . . . .
. . .
. . .
. . .
. . .
. . .
. . .
device
. . .
. . .
. . .
. . .
. . . . .
. . . . .
. . . . .
. . . . .
. . . . .
. . . . .
(SQUID)
. . . . .
. . . . .
. . . . .
. . . . .
165
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
.
173
173
174
174
175
175
175
176
177
177
177
detection methods like an automated or semi-automated DNA, antibody, or biochemical-based method to yield more robust results [5].
1.1.2. Immunology-based methods
This technology relies on immunological based antibodies interaction for the detection of bacterial cells, spores, viruses and toxins alike.
Though the immunological-based detection is not much specic and
sensitive than nucleic acid-based detection, it is faster, more robust
and has the ability to detect not only contaminating organisms but
also their biotoxins that may not be expressed in the organism's genome. Unlike the earlier technique, this technology is being widely
used for the detection of the aforementioned infectious components.
Common examples of immunological techniques include the enzyme
immunoassay (EIA), enzyme linked immunosorbent assay (ELISA),
ow injection immuno-assay, enzyme-linked uorescent assay (ELFA),
bioluminescent enzyme immunoassay (BEIA), enzyme-linked immunomagnetic chemiluminescence (ELIMCL) immunochromatography (ICG)
strip test, immunomagnetic separation, immuno-precipitation assay,
agglutination test, radio-immunoassays (RIA), western blot test and
technically modied western blot include the line immunoassay (LIA)
and the recombinant immunoblot assay (RIBA) [6].
The prerequisite for this technique is detailed understanding of
the inuence of stress on antibody reactions as the physiological activities in cells are often altered in response to a stress. The technique
employs conventional and heavy chain antibodies, as well as polyclonal, monoclonal or recombinant antibodies. In comparison to
monoclonal antibodies, polyclonal antibodies can be raised quickly
and cost effectively however; polyclonal antibodies are limited both
in terms of their specicity and abundance. Hence, monoclonal antibodies are often more useful for specic detection of a wide variety
of microbes and their products because they provide an indenite
supply of single antibody with enhanced sensitivity, specicity, reproducibility and reliability [7].
Despite these salient features, monoclonal antibodies are an expensive alternative to polyclonal antibodies as they require a skilled
technician and specialized growth apparatus for tissue culturing.
Overcoming these issues, are recombinant antibodies since they can
be produced in reasonable quantities in short periods of time from
bacterial expression systems. Limitations of this technique are; inability to detect microorganisms in real-time at low pathogenic load,
low sensitivity of the assays and low afnity of the antibody to the
pathogen or other analyte being measured with potential interference from contaminants.
Likewise, immunology-based methods can be coupled with other
methods for pathogen detection, for instance, immunomagnetic separation on magnetic beads is coupled with matrix-assisted laser desorption ionization-time of ight mass spectrometry for detection of
Staphylococcal enterotoxin B [8], combination of immunomagnetic
separation with ow cytometry for detection of L. monocytogenes [9].
1.1.3. Polymerase chain reaction (PCR)
This method detects a single copy of a target DNA sequence, making
it promising because it detects the organism by amplifying the target
166
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
Fig. 1. Schematic diagram showing main components of Nanodiagnostics.
rather than the signal, and is therefore less prone to producing falsepositives. A target DNA can be amplied 1-million-fold in less than an
hour, with sensitivities in theory down to a single target pathogen.
Also it can be used to enhance the sensitivity of nucleic acid-based assays. PCR has distinct advantages over culture and other standard
methods for the detection of microbial pathogens and offers the advantages of specicity, sensitivity, rapidity, accuracy and capacity to detect
small amounts of target nucleic acid in a sample [10]. To date, PCR based
methods are used in the detection of wide range of pathogens like S. aureus, L. monocytogenes, Salmonella spp., Bacillus cereus, Escherichia coli
O157:H7, Yersinia enterocolitica, and Campylobacter jejuni. [6]. The different PCR based methods used to detect pathogens are real-time PCR,
multiplex PCR and reverse transcriptase PCR (RT-PCR). Compared to
other PCR based methods, multiplex PCR is very useful as it allows the
simultaneous detection of several DNA strands by introducing different
primers to amplify DNA regions coding for specic genes of each bacterial strain targeted. Multiplex PCR assay has been used for rapid and
simultaneous detection of E. coli O157:H7, Salmonella spp., S. aureus,
L. monocytogenes and Vibrio parahaemolyticus [11].
Recently, real-time RT-PCR has emerged as detection technique
which eliminates some purication and concentration steps that are
required for conventional RT-nested PCR detection. Unlike conventional PCR, real-time PCR based technologies are found to be rapid
due to their speed and high degree of sensitivity and specicity, enabling quicker results without too much manipulation. Limitations
of the PCR techniques includes; [7] false negative results due to target
cell lysis necessary for nucleic acid extraction, nucleic acid degradation and/or direct inhibition of PCR. Inability to distinguish between
viable and non-viable cells since DNA is always present whether the
cell is dead or alive. But this limitation can be overcome by using Reverse Transcriptase PCR (RT-PCR). Yaron and Matthews [12] developed an RT-PCR to detect only viable cells of E. coli O157:H7. Also
they suggested that, if RT-PCR is to be used for detection of live cells
in a sample without enrichment, then 10 7 colony-forming units
(cfu) of the target organism are required. Commercially, it can be expensive and complicated, requiring skilled workers to carry out the
tests. PCR is also being coupled with other techniques such as immunouorescent microscopy ELISA, immunomagnetic separation with
the reverse transcription multiplex TaqMan PCR system etc.
1.1.4. Miscellaneous techniques
Other reported analytical methods of pathogen detection include,
application of bacteriophages for detection and control of pathogens,
DNA oligonucleotide arrays, in which specic identication of pathogens was achieved without PCR amplication [6].
2. Potential of nanotechnology
Conventional molecular diagnostic techniques are widely used in
laboratories throughout the world to identify pathogenic agents
with high degree of sensitivity and reproducibility. However, most
of these techniques cannot be utilized in the eld (e.g. airports and
food distribution centers) or in developing countries where resources
are scarce, because they often require sophisticated, expensive instrumentation that needs to be used by trained personnel. Additionally,
the high cost and short shelve half-life of some reagents, such as enzymes and DNA primers, limit the application of most conventional
pathogen detection techniques in rural areas of developing nations.
Furthermore, despite their sensitivity, current technologies, like
ELISA and PCR, require extensive sample preparation and have long
readout times, which delay prompt response and disease containment. Hence, taking advantage of the unique electrical, magnetic, luminescent, and catalytic properties of nanomaterials, faster, sensitive
and more economical diagnostic assays can be developed that can assist in the battle against microbial pathogenesis. Apart from striving
for sensitivity and speed, nanotechnologists have geared their efforts
towards the development of nanotechnology-based systems that are
affordable, robust and reproducible, making them suitable for applications even in rural areas of developing nations. Moreover, using innovative approaches, nanotechnology has the potential to build
assays that can be performed in opaque media, like blood and milk,
without any sample preparation, providing fast and reliable results
in simple and user-friendly formats [5]. Thus this review summarizes
the role of nanotechnology in improvising the existing in-vitro diagnostic testing procedures for accurate and precise diagnosis/detection
of bacterial infections.
Generally diagnostics encompass two components, bioreceptors
and transducers [13]. Briey, the principle of detection involves the
specic binding of the analyte of interest to the complementary biorecognition element (bioreceptors) immobilized on a suitable support
medium. The specic interaction results in a change in one or more
physico-chemical properties which is detected and may be measured
by the transducer. The usual aim is to produce an electronic signal
which is proportional in magnitude or frequency to the concentration
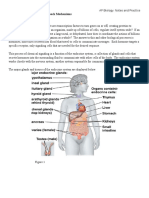
of a specic analyte or group of analytes, to which the biosensing element binds [14,15] (Fig. 1).
3. Nanotechnology based bioreceptors
By denition, bioreceptor is a biological molecular species (e.g. an
antibody, an enzyme, a protein, or a nucleic acid) or a living biological
system (e.g., cells, tissue, or whole organisms) that utilizes a biochemical mechanism for recognition with minimum interference from other
components in complex sampling mixtures. Thus, the primary objective
of the recognition system is to provide the sensor with a high degree of
selectivity for the analyte to be measured. In this perspective, the use of
nanotechnology based bioreceptors could translate into superior sensitivity and may also aid in reduced time of diagnosis. This section highlights the various nanostructures used as in-vitro nanodiagnostic tools
[6] (Table 1).
3.1. Liposomes
Liposomes are small vesicles consisting of one or more concentric
lipid bilayers surrounding aqueous compartments. Particle size and
physicochemical characteristics of liposomes can be manipulated for
specic applications [16].
Gangliosides are expressed on cell surface and serve as natural receptors for bacterial and viral toxins. Therefore, for detection of toxins,
methods have been developed based on gangliosides targeting. Singh
et al. [17] engineered and fabricated GT1b or GM1 ganglioside bearing liposomes (120130 nm) for detecting three bacterial toxins tetanus,
botulinum and cholera. Their study demonstrated that these engineered
liposomes were able to mimic cells and recognize target toxins, thus
forming a basis for toxin detection. These liposomes were labeled with
uorescent markers (rhodamine dyes) followed by a sandwich uoroimmunoassay on antibody coated microtiter plates. Through this
assay, they reported detection of toxins as low as 1 nM concentration.
In a similar study, Durst's group (2003) developed a sensitive bioassay for cholera toxin (CT) detection using GM1 gangliosides containing liposomes ( 200 nm size) [18]. In their sandwich assay, CT
was detected as a colored band on the nitrocellulose membrane
strip, where CT bound to liposomes can be captured by immobilized
antibodies. The limit of detection was found to be 10 fg/ml and the
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
167
Table 1
Summary of nanoparticles studied in the diagnosis of infections.
Pathogen
Nanomaterial
Recognition
Detection
method
Efciency/detection limit
Reference
E. coli
O157:H7
Qdots
Biotinylated
antibody
Antibody
Pathogen
Fluorescence
microscopy
Fluorometry
Visual
2 orders more sensitive than conventional dyes
[57]
100 times more sensitive than FITC
1680 cells
[58]
[59]
Visual detection with minimum detection of
18.75 ng of mycobacterial DNA
Visual detection with 10 cfu/ml of M. tuberculosis
was detected within less than 1 h
Detection without the requirement of any
culturing steps or any other sample pretreatment
techniques
30 times enhance signal amplication as
compared to single Cy3-uorophore-labeled HSV1 DNA
b100 TCID50/ml
[60]
Qdots and magnetic NPs
S. typhimurium Methyl blue (MB) dye entrapped immunoliposomes tagged
with anti-Salmonella common structural antigens (CSA)
antibody
M. tuberculosis Au NPs
Nucleic acid sequence-based amplicationAu NPs
S. aureus
Titanium NPs
MycoD1
UVVis
Oligonucleotide spectroscopy
16S rRNA
UVVis
spectroscopy
Pathogen
MALDI-MS
Human
Dendrimer
Herpes Virus
Oligonucleotide Fluorescent
Inuenza virus Inuenza-specic antibodies conjugated AuNPS
Inuenza virus
Human
Papilloma
Virus
HBV, HCV
antibodies
Qdots
Antibody
Au NPs/Ag staining
Protein A
assay could be completed in 20 min. In another study, Charych's group
(1995) designed sialic acid conjugated liposomes for colorimetric detection of inuenza viruses. These liposomes mimic cell surface molecular recognition and signal transduction. The virus recognition is based
on binding of sialic acid to hemagglutinin lectin present on virus surface. Upon binding to inuenza virus, liposome particles changed
color from blue to pink/orange. This color change was visually detectable and was quantied spectroscopically [19]. Recently, M.R. West
(2009) discussed the synthesis of liposomes consisting of a bacteriasensing structure on the outer surface with a polydiacetylene signal
transducer which gives a dramatic blue-to-red color change in the presence of lipopolysaccarides (LPS) extracted from bacteria cell walls [20].
All these experimental evidences demonstrate that liposomes can
be used for detection of infectious agents and perform equal or even
better than conventional methods.
3.2. Carbon nanotubes
Carbon nanotubes are considered as a sheet of graphite rolled into
a tube with bonds at the end of the sheet that close the tube. A single
walled nanotube (SWNT) can have a diameter of 2 nm and a length of
100 m, making it effectively a one dimensional structure called as
nanowire. The Multiwalled nanotubes (MWNT) can be considered
as SWNTs kept one inside of another. These nanotubes lend themselves for biofunctionaliztion with multiple copies of biomolecules
(e.g. carbohydrates and antibodies) for an enhanced detection of the
antigen in the analyte [21].
Previously, Gu et al. [22] used galactose binding surface proteins
present on pathogenic E. coli. O157:H7 cells to detect the bacteria with
galactose biofunctionalized single walled nanotubes (Gal-SWNTs)
which exhibited efcient aggregation via their multivalent interactions.
While multiwalled carbon nanotubes biofunctionalized with multiple
copies of divalent dendrimeric galactose cluster (Gal2-MWNTs) showed
higher aggregation with detection limit of ~104 cfu/ml (Luo and Stutzenberger, [23]). Elkin et al. [22] also studied the binding interactions
of the goat anti E. coli O157:H7 immunoglobulins adsorbed on a SWNT
already conjugated with Bovin Serum Albumin for detection of the pathogen. Recently, Zelada-Guillen et al. [24] demonstrated the label-free,
rapid detection and identication of living bacteria at concentration
levels of 6 cfu/ml in complex matrices such as milk or 26 cfu/ml in
Dynamic
Light
Scattering
Fluorescence
Protein chip
assay
[61]
[62]
[63]
[64]
100 l sample/1 h/50 times more sensitive
[65]
Visual detection
[66, 67]
apple juice using SWNT based potentiometric aptamer biosensor.
Garca-Aljaro et al. [25] have also shown the utility of SWNT as
rapid immunosensors for the detection of pathogen E. coli O157:
H7 and the bacteriophage T7. The transduction element consisted of
single-walled carbon nanotubes aligned in parallel bridging two gold
electrodes to function as a chemiresistive biosensor. Single-walled carbon nanotubes (SWNTs) were functionalized with specic antibodies
(Ab) for the different microorganisms by covalent immobilization to
the non-covalently bound 1-pyrene butanoic acid succinimidyl ester.
The sensor exhibited a fast response time of 5 min in the case of bacteriophage detection, while the response time for the detection of bacteria was 60 min [25].
3.3. Dendrimers
Dendrimers are hyperbranched, tree-like rigid structures and have
compartmentalized chemical polymers with unique structural and
topological features. Dendrimers generally have a uniform molecular
weight with no specic molecular weight distribution. Dendrimers
contain far more surface groups capable of being functionalized compared with proteins of similar size. In contrast to other linear, crosslinked, and branched polymers, the three-dimensional structure of
dendrimers gives them a variety of unique properties such as low
polydispersity and high functionality, and thus a wide range of potential applications using dendrimers as nanoscopic objects have been
explored [26,27].
During the past two decades a great number of dendrimer structures have been developed and investigated based on inspiration
from biological systems. Among them, polyamidoamine (PAMAM)
dendrimers, polypropyleneimine (PPI) dendrimers, and biomolecules
derived dendrimers such as amino acid dendrimers, carbohydratemodied dendrimers, nucleic acidsnucleobases dendrimers and
polyester dendrimers are notable.
Wang et al.[28] immobilized a 4th generation dendrimer, which
had approximately 30 DNA single-stranded arms specic to the waterborne pathogen Cryptosporidium parvum, onto a quartz-crystal microbalance and employed mass-sensitive piezoelectric transducers to
demonstrate bio-sensibility. When the Cryptosporidium target DNA
accessed the DNA-immobilized dendrimer, a large resonant-frequency
response was obtained due to the resulting three-dimensional surface
168
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
hybridization. Compared with the non-dendritic or unimmobilized
dendrimers, immobilized dendrimers were found to contribute a higher
sensitivity and a wider linear range for biosensing [28].
3.4. Gold nanoparticles (AuNPs)
The unique physical, chemical properties and high surface to volume
ratio enable AuNPs as an ideal material for adsorption of biomolecules
without compromising their biological activities. Antigens or antibodies
functionalized AuNPs can serve as optical labels, electrochemical
markers, surface plasmonic ampliers or signal transfer mediator for
the quantitative analysis of ligands, AuNPs in combination of other signal generators or other nanoparticles could doubly amplify the signal.
They can be prepared with different geometries (size b50 nm) and a
range of shapes are available such as nanospheres, nanoshells, nanorods
or nanocages. The amenability of AuNPs for signal amplication (discussed in detail in Section 4.1.2.1.) makes it a versatile metal nanoparticle for diagnosis [29,5].
Huang [30] used AuNPs for the detection of protein A, which is
found in the cell membrane of most S. aureus strains. This protein
was detected using anti-protein A antibodies functionalized AuNPs
on an immunochromatographic strip. The device took less than
10 min to detect protein A in concentrations of 25 ng/ml [30].
Wang et al. [31] also used AuNPs to functionalize quartz crystal microbalance (QCM) for the detection of DNA from a particular E. coli strain.
Two sizes of AuNPs were used to increase the sensitivity. First, AuNPs of
18 nm diameter were immobilized onto the QCM surface and served as a
platform to support ssDNA that bound specically to the biotinylated
DNA of the target bacteria presenting in the analyte liquid. The use of
this gold layer allowed a greater number of ssDNA molecules to be
bound to the QCM, thus increasing the sensitivity of the sensor. Once
the biotinylated DNA from the target organism had bound to the sensor,
avidin functionalized AuNPs of 70 nm diameter were added in order to
further amplify the signal. This scheme allowed the detection of bacteria
without requiring an enrichment of the sample [31].
An indirect transduction using uorescence was recently
exploited by Philips and co-workers [32], who have reported a rapid
method for detecting bacterial DNA using cationic AuNPs that had
been conjugated with poly(para-phenyleneethynylene) (PPE). In
the bound state, PPE is not uorescent. However, in the presence of
bacteria, electrostatic interactions between bacterial surface and the
AuNPs result in the release of PPE from the conjugate. The level of
free (and therefore uorescent) PPE in a sample is then determined,
allowing for a quick and easy quantication of bacteria [32].
3.5. Conducting polymeric nanoparticles
Since the discovery that conjugated polymers can be made to conduct electricity through doping, conducting polymeric nanoparticles
have become an attractive alternative to silicon nanowires and carbon nanotubes. The conductivity of polymeric NPs is controlled
chemically and have shown to possess unique electrical, electronic,
magnetic and optical properties, apart from its features such as exibility, chemical diversity, more rapid electrochemical switching speed
and ease of processing makes them promising sensing material for
ultra sensitive, trace-level biological and chemical nanosensors. A
thin lm of conducting polymer having both high conductivity, oriented microstructure and the high surface area can prove to be a suitable component for the fabrication of enzyme electrodes and thus can
be used in detection of several analytes with high sensitivity of detection. Moreover, the relative stability is increased due to efcient
bonding of enzyme on the transducer surface which gives it better
reproducibility.
Ease of processability of an organic polymer combined with the
improved mechanical and optical properties of nanoparticles has led
to the fabrication of many devices [33].
Polymeric nanoparticles are used as ultrathin lms, nanowires,
nanotubes or nanocomposites prepared by encapsulation/doping
with labels or biorecognition systems such as electroactive species,
gold nanoparticles, carbon nanotubes or magnetic nanoclusters. The
nanocomposites not only can improve the electrical and mechanical
properties of polymer, but also possess properties of individual components with a synergistic effect. Whenever, there is a contact of these
doped nanowires with the analyte, there are changes in the electrical
characteristics.
Sheng et al. [34] have proposed an ultrasensitive assay for electrical biosensing of syphilis DNA using target-guided formation of polyaniline (PANI) based on an enzymatically catalyzed method. The
sensitive biodetection relies on the DNA hybridization and biotin
streptavidin interaction. After coupling of biotinylated catalase
with streptavidin-modied hybrids, a head-to-tail structure of
PANI templated by hybridized DNA was formed. The current response of PANI was linearly related to target DNA concentration between 1.0 pM and 1.0 nM with a correlation coefcient of 0.998. The
detection limit was determined to be 0.5 pM with the signal-to-noise
ratio (SNR) of 3. The synergistic performances of DNA hybridization,
strong biotinstreptavidin binding ability and highly efcient polymerization provide a general platform for simple, highly sensitive
and selective biosensor for the detection of the specic polA gene
fragment of T. pallidum [34]. Garca-Aljaro et al. [35] have studied
the polypyrrole nanowires (Ppy) assembled onto microfabricated
gold interdigitated microelectrodes, to construct a chemiresistive
biosensor for the detection of Bacillus globigii, used as simulant of
the threatening bioterrorism agent B. anthracis. The fabricated biosensor could detect the spores at the concentrations ranging from 1
to 100 cfu/ml, with a response time of 30 min, after which the sensor
was saturated [35].
Recently, E. B. Setterington et al. [36] isolated E. coli O157:H7 cells
via immunomagnetic separation (IMS) and labeled with biofunctionalized electroactive polyaniline (immuno-PANI). Subsequently, labeled cell complexes were deposited onto a disposable screenprinted carbon electrode (SPCE) sensor and pulled to the electrode
surface by an external magnetic eld, to amplify the electrochemical
signal generated by the polyaniline. The developed biosensor could
detect as low as 7 cfu of E. coli O157:H7 in 70 min (from sampling
to detection), giving it a major advantage over standard culture
methods [36].
3.6. Polystyrene nanoparticles
Polystyrene nanoparticles were used as a carrier for europium
(Eu) III ions that serve as uorescent reporter. Eu (III) has unique
photophysical characteristics (such as sharp line-like emission
peaks, longer lifetime, large stokes shift etc.) that are useful for sensitive detection of biological targets including infectious agents [16].
Hewlett's group developed an immunoassay based on polystyrene
NPs loaded with Eu (III) -diketone chelates for the detection of anthrax
protective agent (PA). The Eu (III) loaded polymeric NPs could be directly integrated to ELISA platform and detection could be performed using
a uorescence plate reader. Their study involved plasma samples spiked
with PA with record dynamic detection range of 0.01 to 100 ng/ml and
the assay sensitivity was approximately 100 fold more than conventional ELISA [37]. A rapid test for malaria diagnosis based on agglutination of sensitive polystyrene particles was developed by
Udomsangpetch et al. [38] to overcome limitations associated with
the high cost of currently available rapid diagnostic tests. This test involves in-vitro aggregation of NPs with antigen or antibody-called
latex-antigen (or antibody) conjugates, in the presence of malaria specic antibody (or antigen). The assay was successfully evaluated for P.
falciparum at an outpatient malaria clinic (Mae Sod, Thailand). Authors
claim the test to be quickest and easiest to be performed in a eld [38].
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
3.7. Iron oxide nanoparticles
Superparamagnetic nanoparticles are being widely used in magnetic immunoassay diagnosis techniques. This approach for magnetic
detection is inspired by the binding effect of the substrates on the
superparamagnetic nanoparticles that can align under an applied
magnetic eld so that uniformly aligned particles are detected by
magnetic detectors; others that are randomly oriented are ignored.
As a result, this technique does not require a washing step before imaging, because other non-specic moieties inside the same such as
buffer or sample will not bind to these particles and thus will not affect the imaging. Bound to a suitable antibody, they are used to label
specic molecules, cell populations and structures or microorganisms.
Binding of antibody to target molecules or disease-causing organism is
used for the immunomagnetic separation of nucleic acids, proteins, viruses, bacteria and cells and it forms the basis of several tests [39].
Superparamagnetic iron oxide nanoparticles/Ferrouids are colloidal solutions (25100 nm in radius) consist of an inorganic core of
iron oxide (magnetite Fe2O3, maghemite or other insoluble ferrites)
coated with polymer to confer stability (such as dextran, polyacrylic
acid or silica), with added functional groups (such as amino and carboxylic acids) to make subsequent conjugations easy. Hence, iron oxide
nanoparticles can carry diverse ligands, such as peptides, small molecules, proteins, antibodies and nucleic acids. Therefore, iron oxide nanoparticles have been used for the identication and quantication of
several targets, including mRNA, DNA, viruses, bacteria and cells. Furthermore, enzymatic and metabolic activities were monitored with
these nanoparticles [40].
Mycobacterium avium spp. paratuberculosis were detected and
quantied in milk and blood through magnetic relaxation [41]. Gu
et.al. [42] used successfully covalently linked vancomycin to highly
magnetic anisotropic FePt magnetic nanoparticles (34 nm) to detect
vancomycin resistant Enterococci and other gram positive bacteria at
concentrations ~ 10 cfu/ml via polyvalent ligand receptor interactions.
Nuria Parera Pera showed the signicance of submicrometer nanoparticles over larger magnetic glycoparticles in the detection of the
zoonotic bacterial pathogen Streptococcus suis. Using submicrometer
magnetic glycoparticles, the presence of adhesion molecule on the
bacteria enabling its binding to the carbohydrate sequence Gal1,4Gal.
Hence, after incubation with various amounts of the pathogen, magnetic concentration and ATP detection, bacterial levels down to 105 cfu
could be detected [43].
3.8. Quantum dots (QDs)
QDs have become one of the most promising and interesting materials for diagnostic applications of bioimaging, labeling, and sensing,
due to their exceptional optical properties. They are nanocrystals
composed of a core of a semiconductor material generally of atoms
from groups II and VI (i.e. CdSe, CdS, and CdTe etc.) or III and V (i.e.
such as InP) of the periodic table, enclosed within a shell of another
semiconductor that has a larger spectral band gap (such as ZnS and
CdS). The shell prevents the surface quenching of excitons in the
emissive core and hence, increase the photostability and quantum
yield for emission. They are neither atomic nor bulk semiconductors,
since their properties originate from their physical size, which ranges
from 10 to 100 in radius. They have high sensitivity, broad excitation spectra, stable-bright uorescence with simple excitation and
no need for lasers. These characteristics make them suitable for various biomedical applications such as sensing and detection of biomarkers including antigens and pathogens, immunolabeling of cells
and tissues. Also their strong light absorbance makes them to be
used as uorescent labels for biomolecules [5,27,44,45].
QDs have been used for rapid and sensitive diagnosis of respiratory syncytial virus (RSV) in a matter of hours allowing detection of the
virus earlier in the course of an infection [46]. When an RSV virus
169
infects lung cells, it leaves part of its coat containing F and G proteins
on the cell's surface. QDs have been linked to antibodies keyed to
structures unique to the RSV coat. As a result, when QDs come in contact with either viral particles or infected cells they stick to their surface. Antibody-conjugated QDs rapidly and sensitively detect RSV and
estimate relative levels of surface protein expression [47]. A major development is the use of dual-color QDs or uorescence energy transfer nanobeads that can be simultaneously excited with a single light
source with improved sensitivity [48].
Thiolated CdSe-core QDs conjugated with wheat germ agglutinin
(WGA), a lectin that is commonly found in gram-positive bacteria,
have shown afnity for bacterial cells, for their ability to bind to sialic
acid and N-acetylglucosaminyl residues present on bacterial cell walls
[49]. QDs can also be bioconjugated with a substrate such as iron, which
is essential for the growth of pathogenic organisms inside a human host.
Pathogenic bacteria normally contain receptors for a human host's own
shuttle protein transferrin, and can harvest iron from transferrin. Using
this metabolism-specic approach, transferrin-conjugated QDs can be
transported through the membrane into metabolically active cells of
iron-deprived S. aureus and detected under uorescence excitation. In
contrast, no QD signal is observed in nonpathogenic bacteria [50]. QDs
have been further conjugated with specic antibodies to detect pathogenic microorganisms such as C. parvum and Giardia lambli [51].
3.9. Bacteriophage/virus particles
Though many methods are available, the low cost and ready production of large numbers of bacteriophage, along with their specicity for the target bacterial species make them ideal for detecting
bacteria. The advantage of detection with bacteriophage/virus particles is that they usually detect living bacteria thereby avoiding the
false positives that often arise from the use of approaches such as
PCR [52,53].
In phage amplication assays, bacterium is infected with the
phage which results in the release of many phage particles that can
be detected by a second (nonpathogenic), sensitive bacterial strain.
Phage based detection systems use engineered recombinant phages,
which upon infection of their hosts, these engineered phage deliver
reporter genes (such as luxAB genes from Vibrio harveyi) [54]. Replication of the viral genome results in many copies of the viral genome
being produced and subsequent expression of these genes ensures the
amplication of the initial phagebacterium interaction into a signal
that can be readily detected (bioluminescence in the case of luxAB) [55].
However, the excess phages, which have not infected the pathogenic bacteria being detected, the sensitivity decreases and hence it
requires that the excess phages to be destroyed.
Multifunctional virus particles conjugated to imaging agents
could serve as a probe for detecting biolm forming S. aureus [16].
Douglas group demonstrated that cowpea chlorotic mottle virus
(CCMV, 28 nm size) could be labeled with uorescent and Magnetic
Resonance Image (MRI) contrast agents. These virus particles were
loaded with imaging contrast agents (uorophore and MRI contrast
agent) and targeted to a biolm of S. aureus bacteria. The targeting
strategy was based on proteinligand interaction that consisted of streptavidin sandwiched between biotinylated CCMV and biotinylated antiSpA monoclonal antibody. The antibody was bound to the protein A
(SpA) that is expressed on the bacterial cell surface. The biotinylated
CCMV was tagged with uorescein, a uorescent dye and the targeting
was conrmed by ow cytometry analysis and epiuorescence microscopy [56].
By means of uorescence imaging, the depth of penetration of the
CCMV particles into S. aureus biolm was observed and the density of
binding was found to be exceptionally high. The CCMV particles were
assessed for its capacity to deliver MRI contrast agent Gadolinium
1,4,7,10-tetraazacyclododecane tetraacetic acid (Gd-DOTA) to S. aureus
cells. The concentration of Gd in the cell was analyzed by inductively
170
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
Fig. 2. Molecular beacon.
coupled plasma mass spectrometry (ICP-MS) which indicated a value of
1.8 105 Gd atoms per bacterial cell. These results demonstrate the usefulness of virus particles for both diagnosis and possible treatment of
biolm forming bacterial infections [16].
4. Nanotechnology based transducers
In principle, signal transduction can be achieved through varied
modes; optical, electrochemical, microgravimetric and magnetic
[6,68,69]. However, for a specic and accurate diagnostic test, the
outcome is governed by the physicochemical properties of the analyte and to an extent on the transducing as well as biorecognition element. Since, diagnosis could entail analysis of infectious analyte in
extremely dilute/ complex media; the design of the overall diagnostic
device becomes very critical. In brief, the transduction element
should enable differential and amplied detection of the target compound without the interference of environmental factors. As widely
known, nanotechnology imparts the unique electrical, magnetic, luminescent, and catalytic properties, which could be translated into efcient transducers, resulting in faster and sensitive diagnostic tools.
Moreover, using innovative approaches, nanotechnology has the
potential to build assays that can be performed in opaque media,
like blood and milk, without any sample preparation, providing fast
and reliable results in simple and user-friendly formats [5].
4.1. Optical biosensors
Optical biosensors have been widely used over the past decade to
analyze biomolecular interactions [such as enzymesubstrate, antigenantibody reactions, DNADNA and peptide nucleic acid (PNA)
DNA hybridizations], providing detailed information on the binding afnity and kinetics of interaction. The advantages of optical biosensors
include; potential sensitivity, readily multiplexed with fast detection,
immunity of the signal to electrical or magnetic interference, adaptability to a wide variety of assay conditions and the potential for higher information content (spectrum of information available), and direct
detection of the target of interest or indirect detection through optically
labeled probes. Overall, the technique is vast and can be broadly classied based on the absorption, reection, refraction, dispersion, infrared,
Raman, chemiluminescence, uorescence, and phosphorescence. However, owing to their sensitivity the most commonly employed techniques include uorescence and surface plasmon resonance [13,6].
4.1.1. Fluorescence-based detection
Fluorescence is a widely used optical method for biosensing due to
its selectivity and sensitivity. Principally, there are three types of biosensing in this method. Direct sensing when a specic molecule is
detected before and after an interaction taking place. It involves measurement of the uorescence quenching when the analyte is near the
surface of the uorophore-labeled optical waveguide. However, binding interactions between an activated signaling molecule and its target could be difcult to detect due to the difculty of seeing this
localized interaction over background uorescence. The use of QDs
as uorophores has overcome most of the drawbacks of using conventional dyes. Indirect biosensing which employs sandwich type approach, where a primary antibody having an afnity for the analyte of
interest is covalently bound to the surface of the ber and the sample
is introduced. Presence of the biomolecule of interest in the sample is
then conrmed using a secondary uorescent antibody, which uoresces when excited by the incident light from the evanescent wave.
Fluorescence biosensing, in which two uorophores are paired in
such a way that the emission wavelength of one overlaps with the excitation wavelength of the other and hence the excitation of one of
them will stimulate uorescence of the complementary pairing one
(if they reside within close proximity of about few angstroms from
each other). It has tremendous utility because the unique uorescence signal generated under these circumstances can be used to visualize and quantify the position and concentration of interacting
uorophores [13,14].
Edgar et al. reported a method that combined in vivo biotinylation
of engineered host-specic bacteriophage and conjugation of the
phage to streptavidin coated QDs. The method provides specic detection of E. coli with as few as 10 cfu/ml in experimental samples
[70].
4.1.1.1. Molecular beacons. They represent a recent technology that
use electronic energy transfer between a uorescent molecule and a
uorescent quencher. The ease of synthesis, unique functionality, molecular specicity and structural tolerance to various modications
has made them an important tool for biomolecular recognition [71].
They are single-stranded oligonucleotide hybridization probes generally 2560 nucleotides in length that form a stem-and-loop structure
and act like switches that are normally closed or off. The loop contains a probe sequence of DNA or RNA that is complementary to a target sequence. The stem structure holds the uorophore and the
quencher in close proximity, preventing the molecular beacons from
emitting uorescence. Binding to the target sequence induces conformational changes that open the loop and as a result the uorescence
is turned on. The resulting spatial separation of the uorophore from
the quencher leads to an enhancement in uorescence signal, because
the non-hybridized molecular beacon has minimal uorescence. The
increasing uorescence signal after each cycle is representative of
the increasing concentration of the amplied sequence (as shown in
Fig. 2).
The majority of surface immobilized probes incorporate a single
dye-based quencher molecule in signal transduction mechanisms.
Most widely used organic quencher, 4-((4-(dimethylamino) phenyl)
azo)benzoic acid (DABCYL), quenches at most 99.0% of the uorescence
of the dye placed in its proximity, but its quenching efciency decreases
for dyes emitting at longer wavelengths [72]. A rst time report for the
presence of Salmonella spp. in fruit and vegetable samples using an oligonucleotide probe that becomes uorescent upon hybridization to the
molecular beacon was evaluated in a real-time PCR assay. As few as
14 cfu per PCR reaction could be detected using the same quencher
(DABCYL) [73]. The studies with a hybrid material composed of a
1.4 nm diameter gold nanoparticle, an organic dye, and a 25nucleotide long ssDNA molecule showed that 1.4 nm diameter gold
nanoparticles can advantageously replace DABCYL as a quencher of
uorescence because they quench uorescence as much as 100 times
better and have higher quenching efciency for dyes emitting near
the infrared region [74].
In human disease diagnostics, molecular beacons immobilized
magnetic nanoparticles, or genomagnetic nanocapturers, have been
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
171
Table 2
Nanoparticles used in optical biosensors.
Nanoparticle (NP)
NP function
QDs composed of CdSe/ZnS core/shell bioconjugates
Signal
E. coli
amplication
Pathogen
LOD
Advantages
106 cells/mL Advantageous with lower detection
thresholds, and better accuracy for
analyzing heterogeneous cell mixtures
Salmonella sequence-specic oligonucleotide with
Biosensing
Salmonella
30 cfu/ml
Simple, rapid and sensitive detection of
2+
the uorescent Ru(bpy) 3-doped silica nanoparticles.
system
typhimurium
bacteria from mixture of bacterial analytes
RSV anti-F protein monoclonal antibody conjugated cadmium Biosensing
Respiratory syncytial
functionalized NPs can provide direct, rapid,
telluride (CdTe)
system
virus strain A2
and sensitive detection of viruses both in-vitro
as well as in vivo
DNA-CdSe/ZnS
Biosensing
Hepatitis B virus
4 nM
Efcient detection of target DNA and
systems
DNA
mutants
PAMAM-OH dendrimer
Signal
P. aeruginosa
350% sensitivity increase compared to that
amplication
without PAMAM-OH
QDs-supported RNA oligonucleotide
Biosensing
Hepatitis C virus
1 ng/ml
Good specicity, ease of performance, and
systems
ability to perform one-spot monitoring
Universal system
Liposome
Signal
RNA sequences from 101000
fmol
amplication B.anthracis, C.
parvum,E. coli
Red quantum dots/novel proprietary phosphorescent
Signal
Reovirus type 3
1:400
Rapid and ultrasensitive detection in
Europium-doped Titanium dioxide nanoparticles
amplication Abney strain
dilution
environmental water samples, food samples,
and fecal samples
developed to collect, separate and detect trace amounts of DNA or
RNA molecules with one single-base difference [75]. Hence, the use
of nanotechnology in combination with molecular beacons seems to
produce real time, superior diagnostic tools.
Although molecular beacons were originally designed to bind and
recognize specic nucleic acids, these probes can also lead to increased uorescence upon binding to certain proteins. The protein
recognition ability of molecular beacons was rst recognized with
an E. coli single-stranded DNA-binding protein. These results demonstrated that while molecular beacons are sensitive and somewhat selective to DNA-binding proteins, they are not specic enough to be
capable of distinguishing a particular protein, and a more selective
protein recognition mechanism is required [76].
References
[81]
[82]
[83]
[84]
[85]
[86]
[87]
[88]
signal-to-noise ratio (SNR). However, unlike gold, the problem with
silver is the low chemical stability [77].
Gold and silver nanoparticles also exhibit localized SPR (LSPR),
wherein the light that interacts with the particles has much smaller
dimension than the incident wavelength, which leads to a plasmon
that oscillates locally around the nanoparticle [78,79]. This phenomenon is largely governed by the chemical nature, size, shape and aggregation of metal nanoparticles. With an increasing nanoparticle size, the
relative contribution of scattering to the extinction increases rapidly.
The strong light scattering from metal nanoparticles enables high contrast optical imaging of biological cells using nanoparticle (including
particles, nanorods, and shells)-labeled conjugates [80] (Table 2).
4.2. Electrochemical biosensors
4.1.2. Refractive index detection
Sometimes labeling of molecules is not desired or possible, and
anyway requires an additional preparation step. The following optical
structures represent the majority of research activities in optical
label-free sensor development: surface plasmon resonance (SPR)
based biosensors; interferometer-based biosensors; resonant mirrors;
and Fiber Bragg gratings. Among these surface plasmon resonance has
gained popularity in recent times.
4.1.2.1. Surface plasmon resonance (SPR) based biosensors [[7] [13],].
Surface plasmon resonance (SPR) effect is a phenomenon involving
the conducting electron oscillations induced by the interaction of
light with a thin metallic lm, typically gold or silver, coated on a
transparent medium. As incident light interacts with the metal interface at angles larger than the critical angle, the reected light displays
a characteristic decrease, due to resonant energy transfer from the incoming photons to surface plasmons. This phenomenon results in the
surface plasmon wave that propagates along the boundary of the
metal lm and the dielectric layer adjacent to it. The wavelength or
angle at which resonance occurs depends on the refractive index of
the analyte and the changes in local refractive index at or near the
metal surface can be used to study the specicity, afnity and kinetics
of biomolecular interactions and to measure the concentration levels
of analytes in complex samples.
Most widely used metals include gold or silver, gold metal enables
increase in the shift of the resonance parameter with respect to
changes in the refractive index thereby increasing the separation between resonance wavelengths consequently increasing sensitivity.
While silver narrows the resonance curve, thereby giving a higher
The electrochemical detection involves induction of the electrochemical property of the analyte by [89]; labeling or tagging one of
the recognition partners with an electroactive species. Commonly
used electrochemical labels could include ferrocene or In 2 + salts,
redox mediators (e.g. K3Fe[(CN)]63/4 ; methylene blue etc.) or enzymes (such as peroxidase, glucose oxidase, alkaline phosphatase or
catalase). Classically, electrochemical techniques suffer limitations
of poor sensitivity and false negative results. Further, there is also a
tendency that the labeling procedure may introduce undesired
changes in the host macromolecules or result in total loss of bioafnity or stability. Moreover, the labeling steps involved in the indirect
techniques can also impose additional time and cost constraints. With
the development of label-free, nanoparticle-modied electrochemical
systems, these limitations have been overcome [See Fig. 3] (Table 3).
Since the modication of the electrode surface by the deposition of
metal nanoparticles leads to the increase in the surface area which
enhances the interaction of biorecognition element with the electrode surface, subsequently resulting in the improved sensitivity [89].
Noble metals like gold have emerged as the most preferred material of construct for nanomodications of biosensors. It possesses inherent redox properties which impart remarkable sensitivity with
detection limits in the pM range [90]. For example, Wang et al. [91]
reported enzyme free approach in which streptavidin and ferrocenyl
hexathiol coated gold nanoparticles were used to monitor the DNA
hybridization by using the ferrocine groups as the reporter molecule
with a linear range for DNA between 7 and 150 pM.
Similarly, modied metal enhanced electrochemical detection approach was explored for detection of two base pair mismatches using
Microcystis as model. Using this methodology, a detection limit of
172
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
Fig. 3. Electrochemical detection of DNA-small molecule detection MED (A) preparation of silver monolayer on gold layer using upd, (B) immobilization of singlestranded probe DNA And (C) detection of target DNA due to complementary base
pairing.
10 pM was obtained for cisplatinshort sequence DNA interactions,
which represents ~100-fold improvement compared with the classical
electrochemical method (1 nM detection limit). [92]. The detection
may also be used to monitor other biomolecular reactions including
DNA hybridization, mismatch detection, DNAprotein, antigenantibody, and DNARNA reactions [93]. Depending on the transduction
mechanism, the electrochemical sensors can use a voltammetric, amperometric, potentiometric, conductometric or impedancemetric detection technique [6].
4.2.1. Anodic stripping voltammetry (ASV) [94]
The ASV is a technique that involves the application of a potential
to an electrode and monitoring of the resulting current owing
through the electrochemical cell. The applied potential forces a
change in the concentration of an electroactive species at the electrode surface by electrochemically reducing or oxidizing it. The subsequent reduction or oxidation results in the mass transport of new
material to the electrode surface and the generation of a current.
Based on this technique, ZnS, CdS, and PdS nanoparticles have been
used in DNA assays for their ability to detect at concentration as
low as 0.3 nM [95,96].
4.2.2. Amperometric sensors
Amperometric biosensors operate by applying a constant potential
and monitoring the current associated with the reduction or oxidation
of an electroactive species involved in the recognition process. The potential of the working electrode is maintained with respect to a reference electrode (usually Ag/AgCl), which is at equilibrium. The most
common working electrodes are noble metals, graphite, modied
forms of carbon or conducting polymers. The current generated is linearly related to the analyte concentration. They are more attractive because of their high sensitivity and wide linear range. However,
amperometric sensors can suffer from poor selectivity, especially
when applying potentials for oxygen/hydrogen peroxide detection,
but this can usually be overcome by using mediators (e.g. iodine, ferrocyanide, etc.) and perm-selective membranes [7,97].
Hasebe et al. [98] described a tyrosinase-based chemically amplied biosensor for the detection of E. coli. They employed a
tyrosinase-coupling electrode to detect polyphenolic compounds
produced microbially from salicyclic acid. The sensor was capable of
detecting 10 310 4 cells/ml but required a 3 h pre-enrichment period.
Brewster and Mazenko [99] also developed an immunoelectrical sensor,
which coupled with ltration capture, allowing rapid detection of E. coli
O157:H7. Cells were incubated with an enzyme-labeled antibody for
15 min and the antibodyantigen (bacteria) complex, captured on a cellulose acetate lter. The lter was brought in contact with the electrode
surface and 5 min after substrate addition, cells were detected by the
conversion of substrate (para-aminophenyl phosphate) to an electroactive product (para-aminophenol). The sensor could detect limit of
5000 cells/ml in an assay time of 25 min [99].
Immunomagnetic beads have also been used to increase the selectivity of amperometric biosensors. This technique requires that
Salmonella typhimurium is sandwiched between antibody coated
magnetic beads and an enzyme (alkaline phosphatase) labeled antibody. A magnet is then used to localize the beads onto the surface of
a disposable graphite ink electrode in a multiwell plate format. Cells
are detected by the oxidation of the electroactive enzyme product.
This offers a LOD of 8 10 3 cells/ml in buffer in a total analysis time
of 80 min [100].
4.2.3. Potentiometric sensors
Potentiometric sensors utilize the measurement of a potential at an
electrode in reference to another electrode. Mostly, it comprises of a
perm-selective outer layer and membrane or sensitive surface to a desired species (a bioactive material), usually an enzyme. The enzymecatalyzed reaction generates or consumes a species, which is detected
by an ion selective electrode. Usually a high impedance voltmeter is
used to measure the electrical potential difference or electromotive
force (EMF) between two electrodes at near zero current. Since potentiometry generates a logarithmic concentration response, the technique
allows the detection of extremely small concentration changes [99].
Table 3
Nanoparticles used in electrochemical biosensors.
Nanoparticle (NP)
NP function
Pathogen
LOD
Poly(vinyl chloride-vinyl acetate maleic acid-)
l-cysteine-gold nanoparticle-CramoLL lectin system
Immobilized CramoLL lectin on a gold electrode
modied with Fe3O4 nanoparticles and PVB
Biosensing
system
Biosensing
system
Bacterial Lipopolysaccharide (LPS)
Advantages
Detection of LPS from bacteria
with glycosyl residues
Reusable and sensitive
Detection of fetuin and glycoproteins
biosensing devices for dengue
from the serum of dengue serotypes 1, 2
infections
and 3
BG antibody self-assembled onto a udp Ag NP deposited Signal
Bacillus globigii
602 spores/ml Lower detection limits
Au electrode
amplication
compared to the conventional
ELISA
Au NP-based graphite epoxy composite
Signal
Salmonella spp. ssDNA
9 fmol
Novel material with improved
amplication
properties
Potassium ferrocyanide encapsulated liposomes
Biosensing
Cholera toxin
1015 g/ml
system
Magnetic NP-coated carbon nanotube nanocomposite Biosensing
E. coli
10 cfu/ml
system
Single-walled carbon nanotubes
Biosensing
S. infantis
100 cfu/ml
Assay time 1 h Label-free
system
system
Chitosan/Fe3O4 nanobiocomposite
Biosensing
HIV
50 pM
system
References
[114]
[115]
[116]
[117]
[118]
[119]
[120]
[121]
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
For enhanced sensitivity and label-free detection, silicon nanowires and carbon nanotubes have been used in fabrication of potentiometric biosensors [101,102]. Recently, light addressable potentiometric sensor (LAPS) has been developed for detection of microbial
contamination [97]. It consists of an n-type silicon semiconductorbased sensor and an insulating layer that is in contact with an aqueous solution where an immuno-reaction takes place. Changes in potential at the silicon-interface are detected by the difference in
charge distribution between the surface of the insulator and the sensor. A LAPS measures an alternating photocurrent generated by a light
source, such as a light emitting diode (LED), so that changes in potential can be transduced into voltage per time differentials. Using this
LAPS technology, molecular devices called Threshold Immunoassay
System has been marketed. In which, an immuno-complex is
formed in the solution phase between sample (antigen), a labeled antibody, biotinylated antibody and streptavidin, which is then captured onto solid phase biotinylated lter. Upon addition of substrate
the lter is brought in contact with the silicon semiconductor and
the enzyme generates a potentiometric signal [7]. This technique
has been exploited for the detection of Y. pestis and B. globigii spores
with a lower LOD of 10 cells or spores per sample [103]. They later
used the same technology to detect S. typhimurium to levels as low
as 119 cfu [104]. Similarly, the technique was used to assay for live
E. coli O157:H7 with a LOD of 2.5 10 4 cells/ml [105]. The US Department of Defense has introduced the Biological Integrated Detection
System for providing detection of airborne biological threats. One of
the instruments incorporated in this is the Biological Detector,
which works on the same principle as the Threshold Immunoassay
System. This system is capable of detecting eight agents simultaneously within 15 min. The LOD for B. subtilis is 3 10 3 cfu/ml [106].
This technique offers potential in the eld of nanodiagnostics although limits of detection still needs some improvement.
4.2.4. Conductometric sensors
The biorecognition event that changes the ionic concentration can
be monitored using conductometric sensors. Normally, they comprise
of two metal electrodes separated by a certain distance and an AC
voltage applied across the electrodes to cause a current ow. The
change in conductance/electrical impedance between the metal electrodes can be measured where the changes can be at an interface or in
the bulk region and can be used to indicate biomolecular reaction between DNA, proteins and antigen/antibody reaction or excretion of
cellular metabolic products.
Conductometric biosensors are characterized by their large sensitivity and represent a new pertinent class of analytical systems for diagnostic applications. Conductance techniques are attractive due to
their simplicity and ease of use since a specialized reference electrode
is not needed and they have been used to detect a wide variety of entities such as agents of biothreat, biochemicals, toxins and nucleic
acids. Conductometric sensors provide information on the ionic
strength in electrolytes and can provide selectivity if coupled with enzyme membranes [6,7,107]. Immobilized nanoparticles functionalized by the anti-coli antibody were shown to have an excellent
sensitivity with detection limit of 1 cfu/ml, coupled with a good specicity of binding [108]
4.2.5. Electrochemical impedance spectroscopy (EIS)
Electrochemical impedance spectroscopy is capable of directly
detecting the specic reaction between a receptor and its ligand. EIS
is widely used to characterize variations in the electronic properties
of bulk materials, and for investigating surface and interfacial processes on electrodes. It is a very efcient technique to measure biological binding events directly at the surface of an electrode. The
chemistry at the surface of the electrode determines the sensitivity
and the specicity of the biosensor. Compared to optical biosensors,
these biosensors offer the advantage of not being affected by the
173
turbidity of the sample, by the uorescence of certain molecules of biological interest, or by the quenching by certain substances [109,110].
Numerous EIS-based bioafnity sensors have been described for
proteins, DNADNA, AbAg or oligonucleotideDNA interactions.
These sensors have been designed by immobilizing bioafnity reagents (such as antigen, antibody, cells, DNA, enzyme layers, and
semiconductors/thin silica layer) at the surface of a solid electrode.
The binding of the molecular recognition partner is then veried
through the detection of either a shift in impedance, or change in capacitance or admittance at the bulk of the electrode interface. The EIS
was utilized for surface characterization of DNA immobilization and
hybridization. [111].
An immunosensor was developed using self-assembled monolayers
(SAM) at the surface of a gold electrode which allowed to detect whole
bacteria (E. coli) with a detection limit of 10 cfu/ml, a considerable improvement against the sensitivity of 107 cfu/ml obtained by surface
plasmon resonance optical detection [112]. The antibodies used in this
EIS biosensor were then coupled to magnetic nanoparticles. This coupling allowed the simultaneous purication of species of interest and
their concentration on the microelectrode by the simple use of a magnet, thus leading to functionalize the microelectrode to detect the corresponding antigen. Measurement of impedance (or admittance) was
used to measure the metabolic activity of microorganisms within
micro-uidic biochips. As bacterial cells are grown within microuidic channels and wells, the impedance changes in the medium can
be detected using electrodes placed appropriately within the channels.
These electrochemical immunosensors were further developed for viral
detection. A microelectrode based impedance immunosensor has been
developed for avian inuenza virus H5N1 detection [113].
4.3. Microgravimetric (Piezoelectrical)
This sensor was introduced in immunoassay format in 1972 by
Shons and associates who coated the quartz crystal surface with an
antigen for the detection of specic antibodies. These are masssensitive biosensors that are based on the detection of mechanical
acoustic waves. They are also referred as Acoustic wave based biosensors operating on the basis of an oscillating crystal that resonates
at a fundamental frequency. After the crystal has been coated with a
biological reagent (such as an antibody) and exposed to the particular
antigen a quantiable change occurs in the resonant frequency of the
crystal, which correlates to mass changes at the crystal surface [6].
In mass amplied piezoelectrical biosensors a sandwich assay format is used, where the sol modied secondary antibody is used. The
large mass of the bound sol particles greatly affects the vibrational
frequency of the quartz crystal and this is used as the basis for detection. The assay can be carried out in the competitive mode. The preferred diameter of sol particles is in the range of 5100 nm [68].
Other high-density particles (e.g. Au, Pt, CdS, TiO2, polymers) may
be also suitable [122,123]. In order to acquire an active surface for
use in a piezoelectric biosensor the surface must be stable chemically,
contain a high number of the actively immobilized biological elements and the coating surface should also be as thin as possible.
The most commonly used piezoelectric materials include quartz
(SiO2) and lithium niobate (LiTaO3) [7,124]. Acoustic wave biosensors
offer label-free, on-line analysis for antigenantibody interactions,
and also provide the option of several immunoassay formats, which
allow increased detection sensitivity and specicity. Other advantages include cost effectiveness combined with ease of use [97]. The
three main types of mass balance acoustic wave transducers are; [6]
4.3.1. Bulk wave (BW) or Quartz crystal microbalance (QCM)
The Bulk wave (BW) device or Quartz crystal microbalance (QCM)
is the oldest and simplest acoustic wave device in operation. This type
of acoustic wave transducers collectively includes bulk acoustic
wave (BAW), quartz crystal resonance sensors (QCRS) and
174
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
Fig. 4. Surface acoustic wave sensor, consisting of a cut crystal with two electrodes on
the same crystal face acting as both transmitter and receiver.
thickness shear mode (TSM) resonator. The difference between
BAW, QCRS and TSM acoustic sensors are their mode of wave propagation [125]. Bulk wave devices consist of parallel circular electrodes
placed on both sides of a thin cut piece of crystal (thin quartz disk in
case of QCM). An alternating electric eld is applied, which induces a
potential difference between the two electrodes causing shear deformation of the crystal. This results in mechanical oscillation of a standing acoustic wave, at a certain characteristic vibrational resonance
frequency across the bulk of the cut quartz. [7].
A piezoelectric quartz crystal microbalance (QCM) nucleic acid
biosensor array using gold nanoparticle signal amplication was developed to rapidly detect S. epidermidis in clinical samples. The synthesized thiolated probes specically targeted towards S. epidermidis
16S rRNA gene were immobilized on the surface of QCM nucleic
acid biosensor arrays. Hybridization was induced by exposing the
immobilized probes to the PCR amplied fragments of S. epidermidis,
resulting in a mass change and a consequent frequency shift of the
QCM biosensor. To further enhance frequency shift results from
above described hybridizations, streptavidin coated gold nanoparticles were conjugated to the PCR amplied fragments. The results
showed the lowest detection limit of current QCM system as
1.3 10 3 cfu/ml [126]. Similarly, Mao et al. [127] developed a QCM
DNA sensor based on nanoparticle-amplication method for detecting E. coli O157:H7. In this study, streptavidin-conjugated Fe3O4
nanoparticles were used as mass enhancers to amplify the frequency
change and the LOD was 2.67 10 2 cfu/ml [127].
In the work by Olsen et al. [128], the use of nanosized phages as
biorecognition elements has shown improved sensitivity. They prepared the biosensors by physically adsorbing the afnity-selected lamentous phage as probes onto piezoelectric transducers for the
rapid detection of S. typhimurium in solution. Specic-bacterial binding resulted in resonance frequency changes of prepared sensors. The
sensors possessed a rapid response time of b180 s, had low-detection
limit of 10 2 cells/ml and were linear over a range of 10 110 7 cells/ml.
Further viscosity effects due to increasing bacterial concentration and
non-specic binding were not signicant as conrmed by doseresponse analysis [128]. Nanduri et al. [129] have developed a biosensor
based on landscape phages immobilized by physical adsorption on
the surface of a QCM for detection of -galactosidase from E. coli
and reported that phage can be used as a recognition element in
biosensors.
4.3.2. Surface acoustic wave (SAW) devices
This type of acoustic wave device transmits along a single crystal
face from one location to another (Fig. 4). In this case the electrodes
are on the same side of the crystal and the transducer can act as
both transmitter and receiver. An excited wave travels across the
crystal face and the physical deformation of the wave is restricted to
the crystal surface. SAW sensors have the ability to directly sense
changes in mass and mechanical properties [7].
SAW have proven to be more sensitive than PQC sensors. However,
numerous problems have been encountered when applying it to a biological sensing system because the surface acoustic wave can become
severely low in biological solutions. [130]. The use of nanostructures
in these sensors have shown to improve the surface acoustic wave sensors. Chou et al. [131] have reported the fabrication of piezoelectric
quartz crystal and surface acoustic wave (SAW) biosensors based on
fullerene C60 and immobilized C60-enzymes/antibodies/proteins for
the detection of various biological species. In this study, the SAW immunosensors were coated with immobilized C60-hemoglobin and C60myoglobin for simultaneous detection of the antibody of proteins, i.e.
anti-hemoglobin and anti-myoglobin respectively in aqueous solutions.
These immunosensors exhibited good sensitivity and excellent detection limit with linear frequency responses to the concentration with
sensitivities of 0.14 and 1.27 kHz g 1 ml 1 respectively. The detection
limits were 0.32 and 0.035 g/ml for anti-Hb and anti-Mb antibodies
respectively.
4.3.3. Magnetoelastic sensors
Recently, magnetoelastic sensor (ME) platform are gaining attention in chemical and biological sensing. ME sensors are constructed
with amorphous ferromagnetic ribbons or wires which are analogous
and complementary to piezoelectric acoustic wave sensors. ME sensors are excited with magnetic AC elds and in turn, they generate
magnetic uxes that can be detected with a sensing coil from a distance and hence these sensors are highly attractive for wireless biosensing. They have high tensile strength and are cost effective. The
fundamental operating principle of the magnetoelastic sensors involves a change in sensor resonance frequency due to mass loading
of the sensor. In biosensing, the change in mass is associated with
the binding of a target analyte to a bioreceptor immobilized on the
surface of the ribbon-like ME sensor. The ME sensor can be coated
with various probe molecules to target analytes. A change in resonant
frequency can be observed if an analyte binds to the magnetoelastic
sensor which can be measured rapidly and accurately [7].
Ruan et al. [132] have used ME sensors to detect Staphylococcal
enterotoxin Type B (SEB). The sensing conguration involved formation of a sandwich antigenantibody complex by the primary afnitypuried anti-SEB antibody covalently immobilized on the surface of a
mass-sensitive magnetoelastic sensor, the target SEB and the biotinlabeled secondary anti-SEB antibody. The study examined the response of the sensor by linearly varying the concentrations of SEB in
the range of 0.55 ng/ml and demonstrated the sensitivity of magnetoelastic sensors. The detection limit was 0.5 ng/ml for 1 h incubation
period. Also the paper reported that, the cost of the sensor was approximately $0.001/sensor, therefore it could be easily utilized as a
disposable sensor.
Ong et al. [133] discussed the fabrication and application of wireless, remote-query ME sensors for the quantication of multiple biological agents. A six-sensor array was fabricated for the simultaneous
measurement of E. coli O157:H7, staphylococcal enterotoxin B and
ricin by immobilizing anti-bacterial or anti-biotoxin antibodies onto
a gold-coated ME sensor through self-assembled monolayer modication cross-linking the antibody with a bifunctional binding agent.
The paper reported that the telemetry of magnetoelastic sensor information is by magnetic ux; hence, no direct connections are needed
between the sensor and monitoring electronic equipment making
possible a variety of in situ and in vivo monitoring applications. Recently, Huang et al. [134] and Xie et al. [135] developed a ME biosensor by
immobilizing bacteriophage as biorecognition element for the realtime in-vitro detection of B. anthracis spores (Table 4).
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
175
Table 4
Nanoparticles used in piezoelectrical (microgravimetric) biosensors.
Nanoparticle (NP)
NP function
Pathogen
LOD
Advantages
References
Oligonucleotide-functionalized AuNP
Biosensing system
Dengue viral RNA
[136]
Antigen coated AuNP
Biosensing system
Anti-T. gondii immunoglobulins
1:5500 dilution
ratio
84 ng/ml
Label-free and increased
senstivity
Direct assay in blood
[138]
[139]
Goat anti-hIgG-coated silica nanoparticles
Signal
Amplication
Mercapto Schistosoma japonicum antigen coated Signal
AuNP
Amplication
Human immunoglobulin G
(hIgG)
Schistosoma japonicum antibody 5 ng/ml
4.4. Magnetic biosensors
Biosensing strategies based on magnetic nanoparticles offer
unique advantages over other techniques as [39,68]; they are inexpensive to produce, physically and chemically stable, biocompatible
and environmentally safe. In addition, biological samples exhibit virtually no magnetic background, and thus highly sensitive measurements can be performed in turbid or otherwise visually obscured
samples without further processing. Although, numerous methods
have been developed to sense biomolecules using magnetic labels;
NMR, SQUID (superconducting quantum interference device) magnetometers and magnetoresistive GMR sensors have been explored
to large extent.
4.4.1. NMR technique
Principally, this technique makes use of magnetic nanoparticles
for detection of biomolecules and cells based on magnetic resonance
effects (also known as diagnostic magnetic resonance, DMR). DMR
assays employ afnity molecule-conjugated magnetic nanoparticles
to bind molecular targets and induce a change in proton relaxation
rate using one of two different procedural modes. The rst involves
tagging large structures such as whole cells, and requires a subsequent wash step to remove unbound nanoparticle sensors (Fig.5).
The second mode makes use of the phenomenon of magnetic relaxation switches (MRSw), in which molecular targets are used to assemble magnetic nanoparticles into clusters and thereby affect a
corresponding change in the bulk relaxation rate. In both cases, the
binding interactions are performed homogeneously in solution and
Fig. 5. Principle of DMR sensing assays using magnetic nanoparticles. a) Tagging cells
with magnetic nanoparticles imparts a magnetic moment that is proportional to the
number of nanoparticles bound. Prior to measurement of the magnetic moment as a
decrease in T2 relaxation time, the unbound nanoparticles are removed from the sample by washing. b) Magnetic relaxation switching assays involve assembly of magnetic
nanoparticle clusters using a target biomarker as a cross-linking bridge, or disassembly
of preformed clusters using an enzyme or competitive binding. Clustering magnetic
nanoparticles causes them to more efciently dephase the nuclear spins of neighboring
water molecules, shortening the transverse relaxation time (T2). Likewise, disassembly
of clusters increases T2 relaxation time.
[137]
make use of a built-in amplication strategy invoked by the magnetic
resonance effect on billions of neighboring water molecules; making
this technique faster than other magnetic detection techniques.
These assays are designed to cause self-assembly of magnetic
nanoparticles upon addition of molecular targets (forward switching,
decreasing T2) [See Fig. 5] or disassembly of preformed clusters by
enzymatic cleavage or competitive binding (reverse switching, increasing T2). Forward MRSw assays are based on crosslinking magnetic
nanoparticles into clusters using molecular target bridges and found
suitable for detecting small molecule analytes such as drugs, metabolites,
oligonucleotides and proteins. Since, the short cross-links ensure that
the magnetic nanoparticles are placed in close enough proximity to promote relaxation switching without the requirement of time-consuming
separation or capture strategies, MRSw assays can be performed in turbid solutions such as blood without the removal of unbound magnetic
nanoparticles. Intact organisms such as viruses and bacteria have successfully been detected using antibody-conjugated MRSw sensors. For
example, adenovirus-5 and herpes simplex virus-1 were detected by
polyclonal antibody-conjugated cross linked iron oxide nanoparticles
in serum with a lower limit of ve viral particles in 10 l [140].
A NMR system is chip-based component used for multiplexed DMR
measurements on smaller sample volumes (about few microliters)
[141]. It consists of microcoils for radio-frequency (RF) excitation and
NMR signal detection, an on-board NMR spectrometer and microuidic
networks. The magnetic eld is generated using a small, portable magnet allowing the analysis of the sample volume to approximately 1 l
with more homogeneous radio-frequency magnetic elds and less electrical resistance. [142]. This technique has been employed for detection
of the bacillus CalmetteGuerin (BCG), a surrogate for Mycobacterium
tuberculosis, standard into sputum samples. The spiked sputum sample
was liqueed by the standard protocol and incubated with magnetic
nanoparticles conjugated with an anti-BCG monoclonal antibody. Unbound magnetic nanoparticles were then removed within the secondgeneration NMR device tted with a porous membrane lter
(~100 nm size cut-off). The membrane allowed passage of the nanoparticles but not BCG, enabling concentration of BCG from larger volumes
of sample and removal of unbound nanoparticles. In this manner, approximately 100 cfu could be detected in 1 l sample volumes using
cross linked iron oxide nanoparticles, and this improved to about 6 cfu
using the more highly magnetic Fe-core/ferrite shell nanoparticles
(cannonballs). Finally, using the membrane lter to concentrate larger
sample volumes, it was demonstrated that as few as 20 cfu could be
detected in a 1 ml sputum sample. This detection threshold is signicantly greater than the AFB smear microscopy technique employed
for TB detection and was comparable to the culture technique but required minutes rather than weeks. [143].
4.4.2. Magnetic eld sensors based on superconducting quantum interference device (SQUID)
The most sensitive of all instruments measuring magnetic eld at
low frequencies (1 Hz) is the superconducting quantum interference
device (SQUID) [144,145]. Typically, the device has three superconducting components: the SQUID ring itself, the radio-frequency coil
and the large antenna loop. The working principle is based on the
176
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
Table 5
Nanoparticles used in magnetic biosensors.
Nanoparticle (NP)
NP function
Pathogen
Magnetic beads
Labeling and
quantication
Y. pestis
Anti-V. parahaemolyticus coated magnetic beads
Biosensing
Magnetic NPs
Signal amplication
Anti-E. coli antibody immobilized amine-functionalized Labeling and isolation
magnetic nanoparticles in combination with adeno- in presence of
external magnetic
sine triphosphate (ATP) bioluminescence
eld
LOD
2.5 ng/ml in
buffer and
human blood
serum
V.
6.0 104 to
parahaemolyticus 5.8 106 cfu/ml
S. aureus
1 103 cfu/ml
E. coli
20 cfu/ml
remarkable interactions of electric currents and magnetic elds; observed when certain materials are cooled below a superconducting
transition temperature. At this temperature, the materials become superconductors and they lose all resistance to the ow of electricity. In
general, the superconducting ring in a SQUID is typically a toroid of
few millimeters in diameter, made up of a metal such as lead or niobium through which a line of magnetic ux is threaded continuously
without any disturbances [142]. The magnitude of the induced current is an exquisitely sensitive indicator of the ux density and magnetic eld intensity [146,147]. It is quantitised by interrupting the
superconducting ring with a weak-link, which is a narrow constriction in the superconductor or a point-contact junction. In SQUID, the
periodic variations are exploited to measure the current in the superconducting ring or the ambient magnetic eld. Typically, a radiofrequency circuit supplies a known bias eld that has been inductively coupled to the ring and also serves as the detector output. Changes
in the ring current alter the resonant frequency of the circuit; as a result, the output signal changes periodically as the eld varies [142].
The third component, i.e. the large antenna loop serves as a magnetic
antenna or dc search coil, effectively gathering ux over an area of
several square centimeters to improve sensitivity. The SQUID ring essentially serves as a very precise ammeter for measuring the current
in the pickup coil. But the sensitivity of SQUIDs is limited by the magnetic eld noise. The power consumption of several watts and bulkiness due to the need of liqueed coolant gasses makes the instrument
less useful in diagnostics and awaits for further modications.
A new technique has been introduced for rapid detection of biological targets by using super-paramagnetic nanoparticles and a microscope based on a high-transition temperature SQUID by Chamala et
al. [141]. In this technique, a mylar lm with bound targets is placed
on the microscope and a suspension of magnetic nanoparticles carrying
antibodies is added to the mixture in a well, and 1 s pulses of magnetic
eld are applied parallel to the SQUID. In the presence of this aligning
eld, the nanoparticles develop a net magnetization, which relaxes
when the eld is turned off. Unbound nanoparticles relax rapidly by
Brownian rotation and contribute no measurable signal. Nanoparticles
bound to the target are captured and undergo Neel relaxation, producing a slowly decaying magnetic ux, which is detected by the SQUID
[148]. The ability to distinguish between bound and unbound labels allows anyone to run homogeneous assays, which do not require separation and removal of unbound magnetic particles [68].
Grossman et al. [149] have employed this technique for detecting
magnetically labeled L. monocytogenes. They incorporated 50-nmdiameter superparamagnetic magnetite particles, coated with antibodies, to an aqueous sample containing L. monocytogenes and applied
a pulsed magnetic eld to align the magnetic dipole moments. The magnetic relaxation signal generated after the eld was turned off and measured using a high-transition temperature SQUID device. Since,
unbound particles randomize direction by brownian rotation, they are
Advantages
References
Off-lab detection unit for medical and
warfare analytes
[157]
Fluorescent sandwich assay with selective [158]
magnetic focusing in presence of other
bacteria. Assay time2 h.
[159]
Allows measurements of turbid samples
without preparation steps. Sample
volumes (510 L). Assay time 15 min
Low cost and reproducible. Assay time
[160]
1 h
not easily detected. In contrast, particles bound to L. monocytogenes
are effectively immobilized and relax in about 1 s by rotation of the
internal dipole moment. Using this method, the detection limit of
(5.6 1.1) 10 6 L. monocytogenes in the sample volume of 20 l
was achieved.
4.4.3. Magnetoresistive-based biosensors
Magnetoresistive based biosensors study the change in the resistivity of a material due to a magnetic eld. Generally, this effect is
exhibited by most of the ferromagnetic alloys (i.e. NiFe, NiFeCo).
Most of the magnetoresistive biosensors are based on giant magneto
resistive (GMR) effect which involves spin dependent interfacial and
bulk scattering asymmetry that is found for spin-up and spin-down
conduction electrons crossing ferromagneticnonmagneticferromagnetic multilayer structures. An applied magnetic eld is used to
change the relative orientation of the magnetisations of the two magnetic layers. When they are aligned, the electrical resistance of the
structure is low while for antiparallel alignment, the resistance is
high [150].
Hence, the detection system relies on the alignment of these moments within the label to produce a measureable fringe eld. Therefore,
a magnetic eld is applied to the chip using a coil or horseshoe electromagnet to induce an overall moment in the labels and also to center the
sensors; i.e. to bias the sensors within the linear regime of their magnetoresistive (magnetic eld) response curve (MR curve). The direction of
the applied eld is dependent on the sensor used, e.g.; in a particular
GMR sensor geometry is either perpendicular to the chip surface or
parallel (in-plane) and at a right angle to the sensor length [151].
The sensor signal obtained (change in voltage) depends on the inherent magnetic sensitivity of the sensor, label: sensor size ratio,
the magnetic moment of the label, the distance between the label
and the sensing layer, the sense current, and application of signal
amplication techniques are used.
Commercially, magnetic nanoparticles product samples contain
particles with heterogeneous size (for example. 200400 nm) and
shape (non-spherical) which hinder in quantication. In addition,
their high resultant magnetisation and anisotropy for their volume
in an applied magnetic eld may lead to rapid clustering (single particles aggregating to form groups). Permanent clustering of labels,
which cannot be remedied using a discriminatory magnetic force
[152] applied to the chip or on-chip washing cycles, can lead to exaggerated positive signals because non-biologically bound labels may
remain attached to biologically bound labels. Consequently, smaller
labels require progressively more sensitive sensors and measurement
systems.
Koets et al. [153] have developed a magneto-resistant sensor
using super-paramagnetic particles as detection labels for E. coli and
Salmonella spp. Magnetic nanoparticles detected by a GMR sensor
were shown to be very powerful and specic. In this work, puried
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
genomic bacterial DNA was used as a template for PCR using a 50-biotin
forward primer and a 50-uorescein reverse primer. The doublestranded-PCR product was then mixed with streptavidin-coated
super-paramagnetic particles. Finally, the particle complexes were captured by anti-uorescein antibodies and detected with the GMR platform [153]. Using this biosensor, 4250 pM amplicon concentrations
were detected in one-step format with total assay times of less than
3 min.
Another example is of immunomagnetic biosensor developed for
label-free detection of E. coli by Mujika et al. [154], they reported a
magneto-resistive immunosensor for the analysis of E. coli O157:H7
in food and clinical samples. This biosensor was capable of detecting
and quantifying small magnetic eld variations caused by the presence of super-paramagnetic beads bound to the antigens previously
immobilized on the sensor surface via an antibodyantigen reaction
[154]. In this same context, Maalouf and co-workers [155] described
an immunomagnetic biosensor for E. coli comprising streptavidinfunctionalized paramagnetic nanobeads attracted to a gold-electrode
surface via a magnetic eld. Biotinylated-anti E. coli antibodies were incubated with nanobeads and detection using impedimetric measurements yielding a working range of 10103 cfu/ml [155].
In recent years, the magnetic properties of some NPs have also
been used for separation or concentration of analytes in biosensing.
Recently, Liebana et al. [156] has reported detection of Salmonella in
skimmed-milk samples by applying immunomagnetic separation/
double-tagging PCR/electrochemical magneto-genosensing with an
LOD of 1 cfu/ml. In this methodology, Salmonella are captured from
milk and preconcentrated by immunomagnetic separation. The bacteria
attached to the magnetic beads are then lysed and the genomic DNA is
thus released. The amplication of the genetic material with a doubletagging set of primers is then performed to conrm the identity of the
bacteria by electrochemical magneto-genosensing [156] (Table 5).
5. Future prospects
Cell-based sensors are an emerging frontier in the area of nanodiagnostics. The use of cells as sensors is a very attractive way to devise sensitive biochemical detectors. With their highly selective and
sensitive receptors, channels and enzymes, intact cells are very attractive candidates for the development of biosensors. The main advantages of the cells as biosensors are that cells have built-in
natural selectivity to biologically active chemicals and they can react
to analytes in a physiologically relevant mode. The transductions of
the cell sensor signals maybe achieved by the measurement of transmembrane and cellular potentials, impedance changes, metabolic activity, analyte inducible emission of genetically engineered reporter
signals, and optically by means of uorescence or luminescence. Signicant challenges exist for long-term operation since the cells need
to be kept alive and healthy under various harsh operating conditions
and much work has been done towards this front, as this technology
has been extended to demonstrate automated portable cell based biosensors platform that have been eld tested [148,161]. Genetically
engineered B cells have been used as sensors, which emit light once
they have been infected by a toxin or a virus [162]. Microorganisms
have also been used as biosensors for the detection and monitoring
of environmental pollutants. Direct measurement of current through
ion channels in the cells has also been used to develop on-chip
patch clamp devices, which can potentially be very sensitive to
changes in the ambient conditions of the cells.
Furthermore, there is also an emergence of new concept Theranostics which very aptly combines the attributes of diagnosis and
therapy to combat infection [163]. Recently, silver nanoparticles a
noble metal known to exhibit strong toxic effects against prokaryotes
with low toxicity against human cells have been tested against a variety
of microbes [reviewed in [164,165]].
177
Kim et al. [167], Sondi and Salopek-Sondi [166] and Fabrega et al.
[168] investigated this activity of silver nanoparticles against E. coli
and other microorganisms In that study, electron microscopy
revealed pits in the cell walls, which is caused by the accumulation
of silver nanoparticles, resulting in alteration of membrane
morphology and increase in permeability, eventually leading to cell
death. Thus, the study very explicitly expounds the role of noble
nanomaterials in curbing the proliferation of bacteria. Hence, the
study provides a new dimension to the role of diagnostic
nanoparticles in diagnosis and thwarting the imminent threat of
bacterial infections [166-168].
Acknowledgement
This work has been supported by the UGC (University Grant Commission). The authors are thankful to Mr. Vinayak Datkhile, and Mr.
Sandeep Jadhav for technical assistance. The authors are also grateful
to Mr. Ramchandra Patale for his valuable suggestions.
References
[1] T. Barker, M. Lesnick, T. Mealey, R. Raimond, S. Walker, D. Rajeski, L. Timberlake,
Nanotechnology and the poor: opportunities and risksclosing the gaps within
and between sectors of society, Jan. 2005www.nanoandthepoor.org Last
accessed 5th Jan. 2010.
[2] Nanomedicine: Nanotechnology for Health, European Technology Platform:
Strategic Research Agenda for Nanomedicine, Nov. 2006, European Commission
[3] C.A. Mirkin, C.S. Thaxton, N.L. Rosi, Nanostructures in biodefense and molecular
diagnostics, Expert. Rev. Mol. Diagn. 4 (6) (2004) 749751.
[4] A.H. Peruski, L.F. Peruski, Immunological methods for detection and identication of infectious disease and biological warfare agents, Clin. Diagn. Lab. Immunol. 10 (4) (2003) 506513.
[5] C. Kaittanis, S.M. Santra, J.M. Perez, Emerging nanotechnology-based strategies
for the identication of microbial pathogenesis, Adv. Drug Deliv. Rev. 62
(2010) 408423.
[6] V. Velusamy, K. Arshak, O. Korostynska, K. Oliwa, C. Adley, An overview of foodborne
pathogen detection: In the perspective of biosensors, Biotechnol. Adv. 28 (2010)
232254.
[7] P. Leonard, S. Hearty, J. Brennan, L. Dunne, J. Quinn, T. Chakraborty, R. O'Kennedy,
Advances in biosensors for detection of pathogens in food and water, Enzyme
Microb. Technol. 32 (2003) 313.
[8] G. Schlosser, P. Kacer, M. Kuzma, Z. Szilagyi, A. Sorrentino, C. Manzo, Coupling
immunomagnetic separation on magnetic beads with matrix-assisted laser desorption ionization-time of ight mass spectrometry for detection of staphylococcal enterotoxin B, Appl. Environ. Microbiol. 73 (2007) 69456952.
[9] K. Hibi, A. Abe, E. Ohashi, K. Mitsubayashi, H. Ushio, T. Hayashi, Combination of
immunomagnetic separation with ow cytometry for detection of Listeria monocytogenes, Anal. Chim. Acta 573 (2006) 158163.
[10] B.C. Millar, J. Xu, J.E. Moore, Molecular diagnostics of medically important bacterial
infections, Mol. Biol. 9 (2007) 2140.
[11] J.S. Kim, G.G. Lee, J.S. Park, Y.H. Jung, H.S. Kwak, S.B. Kim, A novel multiplex PCR
assay for rapid and simultaneous detection of ve pathogenic bacteria: Escherichia
coli O157:H7, Salmonella, Staphylococcus aureus, Listeria monocytogenes, and Vibrio
parahaemolyticus, J. Food Prot. 70 (2007) 16561662.
[12] S. Yaron, K.R. Matthews, A reverse transcriptasepolymerase chain reaction
assay for detection of viable Escherichia coli O157:H7: investigation of specic
target genes, J. Appl. Microbiol. 92 (2002) 633640.
[13] M.N. Velasco-Garcia, Optical biosensors for probing at the cellular level: a review of recent progress and future prospects, Semin. Cell Dev. Biol. 20 (2009)
2733.
[14] T.V. Dinh, Nanosensing at the single cell level, Spectrochim. Acta Part B. 63
(2008) 95103.
[15] T.V. Dinh, P. Kasili, M. Wabuyele, Nanoprobes and nanobiosensors for monitoring
and imaging individual living cells, Nanomedicine 2 (2006) 2230.
[16] P. Tallury, A. Malhotra, L.M. Byrne, S.M. Santra, Nanobioimaging and sensing of
infectious diseases, Adv. Drug Deliv. Rev. 62 (2010) 424437.
[17] A.K. Singh, S.H. Harrison, J.S. Schoeniger, Gangliosides as receptors for biological
toxins: development of sensitive uoroimmunoassays using ganglioside-bearing
liposomes, Anal. Chem. 72 (2000) 60196024.
[18] S. Ahn-Yoon, T.R. DeCory, A.J. Baeumner, R.A. Durst, Gangliosideliposome immunoassay for the ultrasensitive detection of cholera toxin, Anal. Chem. 75
(2003) 22562261.
[19] A. Reichert, J.O. Nagy, W. Spevak, D. Charych, Polydiacetylene liposomes functionalized with sialic acid bind and colorimetrically detect inuenza virus, J. Am.
Chem. Soc. 117 (1995) 829830.
[20] M.R. West, T.W. Hanks, Polydiacetylene-based liposomes: an optical tongue
for bacteria detection and identication, J. Chem. Educ. 86 (2009) 373.
[21] C.P. Poole, F.J. Owens, Chapter 5, Carbon Nanostructures: Introduction to Nanotechnology, A John Wiley & sons, inc., Pub, 2003, pp. 103132.
178
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
[22] L. Gu, T. Elkin, X. Jiang, H. Li, Y. Lin, L. Qu, T.R.J. Tzeng, R. Joseph, Y.P. Sun, Singlewalled carbon nanotubes displaying multivalent ligands for capturing pathogens,
Chem. Commun. (Camb) 21 (2005) 874876.
[23] P.G. Luo, F.J. Stutzenberger, Nanotechnology in the detection & control of microorganisms, Adv. Appl. Microbiol. 63 (2008) 145181.
[24] G.A. Zelada-Guilln, S.V. Bhosale, J. Riu, F.X. Rius, Real-time potentiometric
detection of bacteria in complex samples, Anal. Chem. 82 (22) (2010)
92549260.
[25] C. Garca-Aljaro, L.N. Cella, D.J. Shirale, M. Park, F.J. Muoz, M.V. Yates, A. Mulchandani,
Carbon nanotubes-based chemiresistive biosensors for detection of microorganisms,
Biosens. Bioelectron. 26 (4) (2010) 14371441.
[26] W.D. Janga, K.M.K. Selimb, C.H. Lee, I.K. Kang, Bioinspired application of dendrimers: from bio-mimicry to biomedical applications, Prog. Polym. Sci. 34 (2009)
123.
[27] M. Ozkan, Quantum dots and other nanoparticles: what can they offer to drug
discovery? Drug Discov. Today 9 (24) (2004) 10651071.
[28] J. Wang, M. Jiang, T.W. Nilsen, R.C. Getts, Dendritic nucleic acid probes for DNA
biosensors, J. Am. Chem. Soc. 120 (1998) 82818282.
[29] D. Pissuwan, C.H. Cortie, S.M. Valenzuela, M.B. Cortie, Functionalised gold nanoparticles for controlling pathogenic bacteria, Trends Biotechnol. 4 (28) (2010)
207213.
[30] S.H. Huang, Gold nanoparticle based immunochromatographic assay for the detection of Staphylococcus aureus, Sens. Actuators B 127 (2007) 335340.
[31] L. Wang, Q.-S. Wei, C. Wu, Z.Y. Hu, J. Ji, The Escherichia coli O157:H7 DNA detection on a gold nanoparticle-enhanced piezoelectrical biosensor, Chin. Sci. Bull.
53 (2008) 11571184.
[32] R.L. Philips, O.R. Miranda, C.C. You, V.M. Rotello, U.H.F. Bunz, Rapid and efcient
identication of bacteria using gold nanoparticle-poly(para-phenyleneethynylene) constructs, Angew. Chem. Int. Ed. Engl. 47 (2008) 25902594.
[33] R.T. Ahuja, D. Kumar, Recent progress in the development of nano-structured
conducting polymers/nanocomposites for sensor applications, Sens. Actuators
B 136 (2009) 275286.
[34] Q. Sheng, J. Wang, J. Zheng, Z. Xub, H. Zhang, Ultrasensitive electrical biosensing
of syphilis DNA using target-guided formation of polyaniline based on enzymecatalyzed polymerization, Biosens. Bioelectron. 9 (25) (2010) 20712077.
[35] C. Garca-Aljaroa, M.A. Bangar, E. Baldrich, F.J. Munoz, A. Mulchandan, Conducting
polymer nanowire-based chemiresistive biosensor for the detection of bacterial
spores, Biosens. Bioelectron. 25 (10) (2010) 23092312.
[36] E.B. Setterington, E.C. Alocilja, Rapid electrochemical detection of polyanilinelabeled Escherichia coli O157:H7 Biosens, Bioelectron 26 (5) (2011) 22082214.
[37] S. Tang, M. Moayeri, Z. Chen, H. Harma, J. Zhao, H. Hu, R.H. Purcell, S.H. Leppla,
I.K. Hewlett, Detection of anthrax toxin by an ultrasensitive immunoassay
using Europium nanoparticles, Clin. Vaccine Immunol. 16 (2009) 408413.
[38] D. Polpanich, P. Tangboriboonrat, A. Elaissari, R. Udomsangpetch, Detection of
malaria infection via latex agglutination assay, Anal. Chem. 79 (2007)
46904695.
[39] C. Corot, P. Robert, J.M. Idee, M. Port, Recent advances in iron oxide nanocrystal
technology for medical imaging, Adv. Drug Deliv. Rev. 58 (2006) 14711504.
[40] K.K. Jain, Chapter 3, Nanomolecular diagnostics, The Handbook of Nanomedicine,
Human Press, Totowa, NJ, 2008, pp. 7476.
[41] C. Kaittanis, S.A. Naser, J.M. Perez, One-step, Nanoparticle-mediated bacterial
detection with magnetic relaxation, Nano Lett. 7 (2) (2007) 380383.
[42] H. Gu, P.-L. Ho, K.W.T. Tsang, L. Wang, B. Xu, Using biofunctional magnetic nanoparticles to capture vancomycin resistant enterococci and other gram positive
bacteria at ultra low concentrations, J. Am. Chem. Soc. 125 (51) (2003)
1570215703.
[43] N.P. Pera, A. Kouki, S. Haataja, H.M. Branderhorst, R.M.J. Liskamp, G.M. Visser, J. Finne,
R.J. Pieters, Detection of pathogenic Streptococcus suis bacteria using magnetic glycoparticles, Org. Biomol. Chem. 8 (2010) 24252429.
[44] H.M.E. Azzazy, M.M.H. Mansour, S.C. Kazmierczak, From diagnostics to therapy:
prospects of quantum dots, Clin. Biochem. 40 (2007) 917927.
[45] H.M.E. Azzay, M.M.H. Mansour, S.C. Kazmierczak, Nanodiagnostics: a new frontier
for clinical laboratory medicine, Clin. Chem. 52 (7) (2006) 12381246.
[46] K.K. Jain, Applications of nanobiotechnology in clinical diagnostics, Clin. Chem.
53 (11) (2007) 20022009.
[47] E.L. Bentzen, F. House, T.J. Utley, J.E. Crowe, D.W. Wright, Progression of respiratory
syncytial virus infection monitored by uorescent quantum dot probes, Nano Lett. 5
(2005) 591595.
[48] A. Agarwal, R.A. Tripp, L.J. Anderson, S. Nie, Real time detection of virus particles
and virus protein expression with two color nanoparticle probes, J. Virol. 79
(2005) 86258628.
[49] J.A. Kloepfer, R.E. Mielke, M.S. Wong, K.H. Nealson, G. Stuvky, J.L. Nadeau, Quantum
dots as strain- and metabolism specic microbiological labels, Appl. Environ.
Microbiol. 69 (2003) 42054213.
[50] B. Modun, J. Morrissey, P. Williams, The staphylococcus transferrin receptor: a
glycolytic enzyme with novel functions, Trends Microbiol. 8 (2000) 231237.
[51] L. Zhu, S. Ang, W.T. Liu, Quantum dots as a novel immunouorescent detection
system for Cryptosporium parvum and Giardia lamblia, Appl. Environ. Microbiol.
70 (2004) 597598.
[52] M. Manchester, P. Singh, Virus-based nanoparticles (VNPs): platform technologies
for diagnostic imaging, Adv. Drug Deliv. Rev. 58 (2006) 15051522.
[53] N.k. Petty, T.J. Evans, P.C. Fineran, G.P.C. Salmond, Biotechnological exploitation
of bacteriophage research, Trends Biotechnol. 1 (25) (2006) 715.
[54] M. Oda, M. Morita, H. Unno, Y. Tanji, Rapid detection of Escherichia coli O157:H7
by using green uorescent protein labeled PP01 bacteriophage, Appl. Environ.
Microbiol. 70 (2004) 527534.
[55] M.J. Loessner, C.E. Rees, G.S. Stewart, S. Scherer, Construction of luciferase reporter of bacteriophage A511: luxAB for rapid and sensitive detection of viable
listaria cells, Appl. Environ. Microbiol. 62 (1996) 11331140.
[56] P.A. Suci, D.L. Berglund, L. Liepold, S. Brumeld, B. Pitts, W. Davison, L. Oltrogge,
K.O. Hoyt, S. Codd, P.S. Stewart, M. Young, T. Douglas, High-density targeting of a
viral multifunctional nanoplatform to a pathogenic biolm-forming bacterium,
Chem. Biol. 14 (2007) 387398.
[57] M.A. Hahn, J.S. Tabb, T.D. Krauss, Detection of single bacterial pathogens with
semiconductor quantum dots, Anal. Chem. 77 (2005) 48614869.
[58] X.L. Su, Y. Li, Quantum dot biolabeling coupled with immunomagnetic separation for detection of Escherichia coli O157: H7, Anal. Chem. 76 (2004)
48064810.
[59] J.A. Ho, S.C. Zheng, W.H. Tseng, Y.J. Lin, C.H. Chen, Liposome-based immunostrip
for the rapid detection of Salmonella, Anal. Bioanal. Chem. 391 (2008) 479485.
[60] E. Liandris, M. Gazouli, M. Andreadou, M. Comor, N. Abazovic, L.A. Sechi, J. Ikonomopoulos, Direct detection of unamplied DNA from pathogenic mycobacteria
using DNA-derivatized gold nanoparticles, J. Microbiol. Methods 78 (2009)
260264.
[61] P. Gill, M. Ghalami, A. Ghaemi, N. Mosavari, H. Abdul-Tehrani, M. Sadeghizadeh,
Nanodiagnostic method for colorimetric detection of Mycobacterium tuberculosis
16S rRNA, Nanobiotechnol 4 (2008) 2835.
[62] J. Gopal, J.L. Narayana, H.F. Wu, TiO2 nanoparticle assisted mass spectrometry as
biosensor of Staphylococcus aureus, key pathogen in nosocomial infections from
air, skin surface and human nasal passage, Biosens. Bioelectron. 27 (1) (2011)
201206.
[63] H.M. Striebel, E. Birch-Hirschfeld, R. Egerer, Z. Foldes-Papp, G.P. Tilz, A. Stelzner,
Enhancing sensitivity of human herpes virus diagnosis with DNA microarrays
using dendrimers, Exp. Mol. Pathol. 77 (2) (2004) 8997.
[64] J.D. Driskell, C.A. Jones, S.M. Tompkins, R.A. Tripp, One-step assay for detecting
inuenza virus using dynamic light scattering and gold nanoparticles, Analyst
136 (15) (2011) 30833090.
[65] J.M. Klostranec, Q. Xiang, G.A. Farcas, J.A. Lee, A. Rhee, E.I. Lafferty, S.D. Perrault,
K.C. Kain, W.C.W. Chan, Convergence of quantum dot barcodes with microuidics and signal processing for multiplexed high-throughput infectious disease
diagnostics, Nano Lett. 7 (2007) 28122818.
[66] Y.F. Wang, D.W. Pang, Z.L. Zhang, H.Z. Zheng, J.P. Cao, J.T. Shen, Visual gene diagnosis
of HBV and HCV based on nanoparticle probe amplication and silver staining enhancement, J. Med. Virol. 70 (2003) 205211.
[67] L. Duan, Y. Wang, S.S.C. Li, Z. Wan, J. Zhai, Rapid and simultaneous detection of
human hepatitis B virus and hepatitis C virus antibodies based on a protein chip
assay using nano-gold immunological amplication and silver staining method,
BMC. Infect. Dis. 5 (53) (2005).
[68] N. Sanvicens, C. Pastells, N. Pascual, M.P. Marco, Nanoparticle-based biosensors
for detection of pathogenic bacteria, Trends Anal. Chem. 28 (2009) 12431251.
[69] C. Jianrong, M. Yuqing, H. Nongyue, W. Xiaohua, L. Sijiao, Nanotechnology and
biosensors, Biotechnol. Adv. 22 (2004) 505518.
[70] R. Edgar, M. McKinstry, J. Hwang, A.B. Oppenheim, R.A. Fekete, G. Giulian, C.
Merril, K. Nagashima, S. Adhya, High sensitivity bacterial detection using
biotin-tagged phage and quantum-dot nanocomplexes, Proc. Natl. Acad. Sci. U.
S. A. 13 (103) (2006) 48414845.
[71] W. Tan, K. Wang, T.J. Drake, Molecular beacons, Curr. Opin. Chem. Biol. 8 (2004)
547553.
[72] S. Tyagi, F.R. Kramer, Molecular beacons: probes that uoresce upon hybridization, Nat. Biotechnol. 14 (1996) 303308.
[73] S.H. Liming, A. Bhagwat, Application of a molecular beaconreal-time PCR technology to detect Salmonella species contaminating fruits and vegetables, Int. J.
Food Microbiol. 95 (2004) 177187.
[74] B. Dubertret, M. Calame, A.J. Libchaber, Single-mismatch detection using goldquenched uorescent oligonucleotides, Nat. Biotechnol. 19 (4) (2001) 365370.
[75] X. Zhao, R. Tapec-Dytioco, K. Wang, W. Tan, Collection of trace amounts of
DNA/mRNA molecules using genomagnetic nanocapturers, Anal. Chem. 75
(2003) 34763483.
[76] J. Li, X. Fang, S. Schuster, W. Tan, Molecular beacons: a novel approach to detect
proteinDNA interactions, Angew. Chem. Int. Ed. 6 (39) (2000) 10491052.
[77] A. Leung, P.M. Shankar, R. Mutharasan, A review of ber optic biosensors, Sens.
Actuators B 125 (2007) 688703.
[78] A.J. Haes, R.P. Van Duyne, Nanoscale optical biosensors based on localized surface plasmon resonance spectroscopy, Proc. SPIE 5221 (2003) 4758.
[79] M.C. Daniel, D. Astruc, Gold nanoparticles: assembly, supramolecular chemistry,
quantum size-related properties, and applications toward biology, catalysis, and
nanotechnology, Chem. Rev. 104 (2004) 293346.
[80] W.P. McConnell, J.P. Novak, L.C. Brousseau III, R.R. Fuierer, R.C. Tenent, D.L. Feldheim, Electronic and optical properties of chemically modied metal nanoparticles and molecularly bridged nanoparticle arrays, J. Phys. Chem. B 104 (2000)
89258930.
[81] M.A. Hahn, P.C. Keng, T.D. Krauss, Flow cytometric analysis to detect pathogens
in bacterial cell mixtures using semiconductor quantum dots, Anal. Chem. 3 (80)
(2008).
[82] Z. Wang, H. Xu, J. Wu, J. Ye, Z. Yang, Sensitive detection of Salmonella with uorescent bioconjugated nanoparticles probe, Food Chem. 125 (2011) 779784.
[83] R.A. Tripp, R. Alvarez, B. Anderson, L. Jones, C. Weeks, W. Chen, Bioconjugated
nanoparticle detection of respiratory syncytial virus infection, Int. J. Nanomedicine 2 (1) (2007) 117124.
[84] X. Wang, X. Lou, Y. Wang, Q. Guo, Z. Fang, X. Zhong, H. Mao, Q. Jin, L. Wu, H.
Zhao, J. Zhao, QDs-DNA nanosensor for the detection of hepatitis B virus DNA
and the single-base mutants, Biosens. Bioelectron. 25 (8) (2010) 19341940.
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
[85] J. Ji, J.A. Schanzle, M.B. Tabacco, Real-time detection of bacterial contamination
in dynamic aqueous environments using optical sensors, Anal. Chem. 76
(2004) 14111418.
[86] C. Roh, H.Y. Lee, S.E. Kim, S.K. Jo, A highly sensitive and selective viral protein detection method based on RNA oligonucleotide nanoparticle, Int. J. Nanomedicine
5 (2010) 323329.
[87] C. Fournier-Wirth, J. Coste, Nanotechnologies for pathogen detection: future alternatives? Biologicals 38 (2010) 913.
[88] J.G. Bruno, K. Francis, Ikanovic Milada, P. Rao, S. Dwarakanath, W.E. Rudzinski,
Reovirus detection using immunomagnetic-uorescent nanoparticle sandwich
assays, J. Bionanosci. 1 (2007) 16.
[89] O.A. Sadik, A.O. Aluoch, A. Zhou, Status of biomolecular recognition using electrochemical techniques, Biosens. Bioelectron. 24 (2009) 27492765.
[90] J. Wang, R. Polsky, A. Merkoci, K.L. Turner, Electroactive beads for ultrasensitive
DNA detection, Langmuir 19 (4) (2003) 989991.
[91] J. Wang, J. Li, A.J. Baca, J. Hu, F. Zhou, W. Yan, D.W. Pang, Amplied voltammetric detection of DNA hybridization via oxidation of ferrocene caps on gold nanoparticle/streptavidin conjugates, Anal. Chem. 75 (15) (2003) 39413945.
[92] I.O. K'Owino, S.K. Mwilu, O.A. Sadik, Metal-enhanced biosensor for genetic mismatch detection, Anal. Biochem. 369 (2007) 817.
[93] A.O. Aluoch, O.A. Sadik, G. Bedi, Development of an oral biosensor for salivary
amylase using a monodispersed silver for signal amplication, Anal. Biochem.
340 (1) (2005) 136144.
[94] S. P. Kounaves, In: F. A. Settle (Ed.) Chapter 37. Voltammetric techniques, A.,
Handbook of Instrumental Techniques for Analytical Chemistry, Prentice hall
PTR, upper saddle river, NJ, 1997.
[95] J. Wang, G.D. Liu, G. Rivas, Encoded beads for electrochemical identication,
Anal. Chem. 75 (17) (2003) 46674671.
[96] J. Wang, D. Xu, R. Polsky, Magnetically-induced solid-state electrochemical detection of DNA hybridization, J. Am. Chem. Soc. 124 (2002) 42084209.
[97] D. Ivnitski, I. Abdel-Hamid, P. Atanasov, E. Wilkins, Biosensors for the detection
of pathogenic bacteria, Biosens. Bioelectron. 14 (1999) 599624.
[98] Y. Hasebe, K. Yokobari, K. Fukasawa, T. Koguru, S. Uchiyama, Highly sensitive
electrochemical determination of Escherichia coli density using tyrosinasebased chemically amplied biosensor, Anal. Chim. Acta 357 (1997) 5154.
[99] J.D. Brewster, R.S. Mazenko, Filtration capture and immunoelectrochemical detection for rapid assay of Escherichia coli O157:H7, J. Immunol. Methods 211
(1998) 18.
[100] A.G. Gehring, C.G. Crowford, R.S. Mazenko, L.J. Van Houten, J.D. Brewster, Enzyme-linked immunomagnetic electrochemical detection of salmonella typhimurium, J. Immunol. Methods 195 (1998) 1525.
[101] Y. Cui, Q. Wei, H. Park, C.M. Lieber, Nanowire nanosensors for highly sensitive
and selective detection of biological and chemical species, Science 293 (2001)
12891292.
[102] K. Besteman, J.L. Lee, F.G.M. Wiertz, H.A. Heering, C. Dekkar, Enzyme-coated carbon
nanotubes as single-molecule biosensors, Nano Lett. 3 (2003) 727730 Nano Lett. 3
(2003) 727.
[103] K. Dill, J.H. Song, J.A. Blomdahl, J.D. Olson, Rapid, sensitive and specic detection
of whole cells and spores using the light addressable potentiometric sensor, J.
Biochem. Biophys. Methods 34 (1997) 161166.
[104] K. Dill, L.H. Stanker, C.R. Young, Detection of salmonella in poultry using a silicon
chip-based biosensor, J. Biochem. Biophys. Methods 41 (1999) 6167.
[105] A.G. Gehring, D.L. Paterson, S. Tu, Use of a light addressable potentiometric sensor for the detection of Escherichia coli O157:H7, Anal. Biochem. 258 (1998)
293298.
[106] K.A. Uithoven, J.C. Schmidt, M.E. Ballman, Rapid identication of biological warfare agents using an instrument employing a light addressable-potentiometric
sensor and a ow-through immunoltration-enzyme assay system, Biosens.
Bioelectron. 14 (2000) 761770.
[107] R. Bashir, BioMEMS: state-of-the art in detection, opportunities and prospects,
Adv. Drug Deliv. Rev. 56 (2004) 15651586.
[108] M. Hnaien, W.M. Hassen, A. Abdelghani, C. Fournier-Wirth, J. Coste, F. Bessueille,
A conductometric immunosensor based on functionalized magnetite nanoparticles for E. Coli detection, Electrochem Commun 10 (2008) 11521154.
[109] Chapter 14, in: E. Barsoukov, J. MacDonald (Eds.), Impedence Spectroscopy: Theory, Experiment and Applications, 2nd ed, John Wiley and Sons, Inc, Hoboken, New
Jersey, 2005.
[110] D.D. MacDonald, Application of electrochemical impedance spectroscopy in
electrochemistry and corrosion science, in: R. Varma, J.R. Selman (Eds.), Techniques for Characterization of Electrodes and Electrochemical Process, John
Wiley & Sons, New York, 1991, pp. 515580.
[111] N. Jaffrezic-Renault, C. Martelet, Y. Chevolot, J.P. Cloarec, Biosensors and biobarcode
assays based on biofunctionalised magnetic microbeads, Sensor 7 (2007) 589614.
[112] R. Maalouf, C. Fournier-Wirth, J. Coste, H. Chebib, Y. Saikali, O. Vittori, Label free
detection of bacteria by electrochemical impedence spectroscopy: comparison
of surface plasmon resonance, Anal. Chem. 79 (2007) 48794886.
[113] R. Wang, Y. Wang, K. Lassiter, Y. Li, B. Hargis, S. Tung, L. Berghmand, W. Bottje,
Interdigitated array microelectrode based impedance immunosensor for detection of avian inuenza virus H5N1, Talanta 79 (2009) 159164.
[114] M.D.L. Oliveria, C.A.S. Andrade, M.T.S. Correia, L.C.B.B. Coelho, P.R. Singh, X. Zeng,
Impedimetric biosensor based on self-assembled hybrid cystein-gold nanoparticles and CramoLL lectin for bacterial lipopolysaccharide recognition, J. Colloid
Interface Sci. 362 (2011) 194201.
[115] M.D.L. Oliveria, M.L. Nogueira, M.T.S. Correia, L.C.B.B. Coelho, C.A.S. Andrade, Detection of dengue virus serotypes on the surface of gold electrode based on Cratylia
mollis lectin afnity, Sens. Actuators, B 155 (2011) 789795.
179
[116] S.K. Mwilu, A.O. Aluoch, S. Miller, P. Wong, O.A. Sadik, Identication and quantitation of Bacillus globigii using metal enhanced electrochemical detection and
capillary biosensor, Anal. Chem. 81 (18) (2009) 75617570.
[117] P.R. Marques, A. Lermo, S. Campoy, H. Yamanaka, J. Barbe, S. Alegret, M.I. Pividori, Double-tagging polymerase chain reaction with a thiolated primer and
electrochemical genosensing based on gold nanocomposite sensor for food safety, Anal. Chem. 81 (2009) 13321339.
[118] S. Viswanathan, L.C. Wu, M.R. Huang, J.A. Ho, Electrochemical immunosensor for
cholera toxin using liposomes and poly(3,4-ethylenedioxythiophene)-coated
carbon nanotubes, Anal. Chem. 78 (2006) 11151121.
[119] Y. Cheng, Y. Liu, J. Huang, K. Li, Y. Xian, W. Zhang, L. Jin, Amperometric tyrosinase
biosensor based on Fe3O4 nanoparticles-coated carbon nanotubes nanocomposite for rapid detection of coliforms, Electrochim. Acta 54 (9) (2009) 25882594.
[120] R.A. Villamizar, A. Maroto, F.X. Rius, I. Inza, M.J. Figueras, Fast detection of Salmonella
infantis with carbon nanotube eld effect transistors, Biosens. Bioelectron. 24 (2008)
279283.
[121] L.D. Tran, B.H. Nguyen, N.V. Hieu, H.V. Tran, H.L. Nguyen, P.X. Nguyen, Electrochemical detection of short HIV sequences on chitosan/Fe3O4 nanoparticle
based screen printed electrodes, Mater. Sci. Eng., C 31 (2011) 477485.
[122] X. Su, F.T. Chew, S.F.Y. Li, Design and application of piezoelectric quartz crystalbased immunoassay, Anal. Sci. 16 (2) (2000) 107114.
[123] T. Liu, J. Tang, L. Jiang, The enhancement effect of gold nanoparticles as a surface
modier on DNA sensor sensitivity, Biochem. Biophys. Res. Commun. 313
(2004) 37.
[124] P.D. Skottrup, M. Nicolaisen, A.F. Justesen, Towards on-site pathogen detection
using antibody-based sensors, Biosens. Bioelectron. 24 (2008) 339348.
[125] M.A. Cooper, V.T. Singleton, A survey of the 2001 to 2005 quartz crystal microbalance biosensor literature: applications of acoustic physics to the analysis of
biomolecular interactions, J. Mol. Recognit. 20 (2007) 154184.
[126] H. Xia, F. Wang, Q. Huang, J. Huang, M. Chen, J. Wang, C. Yao, Q. Chen, G. Cai, W.
Fu, Detection of Staphylococcus epidermidis by a quartz crystal microbalance
nucleic acid biosensor array using Au nanoparticle signal amplication, Sensors
8 (2008) 64536470.
[127] X. Mao, L. Yang, X.-L. Su, Y. Li, A nanoparticle amplication based quartz crystal
microbalance DNA sensor for detection of Escherichia coli O157:H7, Biosens.
Bioelectron. 21 (2006) 11781185.
[128] E.V. Olsen, I.B. Sorokulova, V.A. Petrenko, I.H. Chen, J.M. Barbaree, V.J. Vodyanoy,
Afnity-selected lamentous bacteriophage as a probe for acoustic wave biodetectors of Salmonella typhimurium, Biosens. Bioelectron. 21 (8) (2006)
14341442.
[129] V. Nanduri, I.B. Sorokulova, A.M. Samoylov, A.L. Simonian, V.A. Petrenko, V.
Vodyanoy, Phage as a molecular recognition element in biosensors immobilized
by physical adsorption, Biosens. Bioelectron. 22 (2007) 986992.
[130] R.L. Bunde, E.J. Jarvi, J.J. Rosentreter, Piezoelectric quartz crystal biosensors,
Talanta 46 (1998) 12231236.
[131] F.F. Chou, H.W. Chang, T.L. Li, J.S. Shih, Piezoelectric crystal/surface acoustic
wave biosensors based on fullerene C60 and enzymes/antibodies/proteins, J.
Iran. Chem. Soc. 5 (1) (2008) 115.
[132] C.M. Ruan, K.F. Zeng, O.K. Varghese, C.A. Grimes, A staphylococcal enterotoxin B
magnetoelastic immunosensor, Biosens. Bioelectron. 20 (2004) 585591.
[133] K.G. Ong, K. Zeng, X. Yang, K. Shankar, C. Ruan, C.A. Grimes, Quantication of
multiple bioagents with wireless, remote-query magnetoelastic microsensors,
IEEE Sens. J. 6 (2006).
[134] S. Huang, S.-Q. Li, H. Yang, M. Johnson, J. Wan, I. Chen, Optimization of phage-based
magnetoelastic biosensor performance, Sens. Transducers 3 (2008) 8796.
[135] F. Xie, H. Yang, S. Li, W. Shen, J. Wan, M.L. Johnson, Amorphous magnetoelastic
sensors for the detection of biological agents, Intermetallics 17 (2009) 270273.
[136] S.H. Chen, Y.C. Chuang, Y.C. Lu, H.C. Lin, Y.L. Yang, C.S. Lin, A method of layer-bylayer gold nanoparticle hybridization in a quartz crystal microbalance DNA sensing
system used to detect dengue virus, Nanotechnol 20 (2009) 215501215511.
[137] H. Wang, C.X. Lei, J.S. Li, Z.Y. Wu, G.L. Shen, R.Q. Yu, A piezoelectric immunoagglutination assay for Toxoplasma gondii antibodies using gold nanoparticles, Biosens.
Bioelectron. 19 (2004) 701709.
[138] X.-Y. Jin, X.-F. Jin, Y.-J. Ding, J.-H. Jiang, G.-L. Shen, R.-Q. Yu, A piezoelectric
immunosensor based on agglutination reaction with amplication of silica
nanoparticles, Chin. J. Chem. 26 (12) (2008) 21912196.
[139] W. Zhaoyang, W. Jian, S. Wang, G. Shen, R. Yu, An amplied mass piezoelectric
immunosensor for Schistosoma japonicum, Biosens. Bioelectron. 22 (2006)
207212.
[140] J.M. Perez, F.J. Simeone, Y. Saeki, L. Josephson, R. Weissleder, Viral-induced selfassembly of magnetic nanoparticles allows the detection of viral particles in biological
media, J. Am. Chem. Soc. 125 (2003) 1019210193.
[141] Y.R. Chemla, H.L. Grossman, Y. Poon, R. McDermott, R. Stevens, M.D. Alpert,
Ultrasensitive magnetic biosensors for homogeneous immunoassay, Proc. Natl.
Acad. Sci. U. S. A. 97 (2000) 1426814272.
[142] J. Lenz, A.S. Edelstein, Magnetic sensors and their applications, IEEE Sens. J. 6 (3)
(2006) 631649.
[143] H. Lee, T.J. Yoon, R. Weissleder, Ultrasensitive detection of bacteria using coreshell nanoparticles and an NMR-lter system, Angew. Chem. Int. Ed. 48 (2009)
56575660.
[144] V. Pizzella, S.D. Penna, C.D. Gratta, G.L. Romani, SQUID systems for biomagnetic
imaging, Supercond. Sci. Technol. 14 (2001) R79R114.
[145] R. Cantor, SQUID's and emerging applications, Supercond. Cryoelectron 13
(2001) 1622.
[146] B.S. Deaver, W.M. Fairbank, Experimental evidence for quantized ux in superconducting cyclinders, Phys. Rev. Lett. 7 (1961) 4346.
180
S.B. Shinde et al. / Journal of Controlled Release 159 (2012) 164180
[147] R. Doll, M. Nabauer, Experimental proof of magnetic ux quantization in a
superconducting ring, Phys. Rev. Lett. 7 (1961) 5152.
[148] L. Bousse, Whole cell biosensors, Sens. Actuators, B 34 (13) (1996) 270275.
[149] H.L. Grossman, W.R. Myers, V.J. Vreeland, R. Bruehl, M.D. Alper, C.R. Bertozzi, J.
Clarke, Detection of bacteria in suspension by using a superconducting quantum
interference device, Proc. Natl. Acad. Sci. U. S. A. 101 (1) (2004) 129134.
[150] D.L. Graham, H.A. Ferreira, P.P. Freitas, Magnetoresistive-based biosensors and
biochips, Trends Biotechnol. 9 (22) (2004) 455462.
[151] U. Hartman (Ed.), Magnetic thin lm and multilayer systems: physics, analysis
and industrial applications, Springer Series in Mater. Sci., , 1996.
[152] G.U. Lee, S. Metzgerc, M. Natesanc, C. Yanavichd, Y.F. Dufrene, Implementation
of force differentiation in the immunoassay, Anal. Biochem. 287 (2000)
261271.
[153] M. Koets, T. van der Wijk, J.T.W.M. van Eemeren, A. van Amerongen, M.W.J.
Prins, Rapid DNA multi-analyte immunoassay on a magneto-resistance biosensor,
Biosens. Bioelectron. 24 (2009) 18931898.
[154] M. Mujika, S. Arana, E. Castano, M. Tijero, R. Vilares, J.M. Ruano-Lopez, A. Cruz, L.
Sainz, J. Berganza, Magnetoresistive immunosensor for the detection of Escherichia
coli O157:H7 including a microuidic network, Biosens. Bioelectron. 24 (2009)
12531258.
[155] R. Maalouf, W.M. Hassen, C. Fournier-Wirth, J. Coste, N. Jaffrezic-Renault, Comparison of two innovative approaches for bacteria detection: paramagnetic
nanoparticle and self assembles multilayer processes, Microchim. Acta 163
(2008) 157161.
[156] S. Liebana, A. Lermo, S. Campoy, J. Barbe, S. Alegret, M.I. Pividori, Magneto Immunoseparation of pathogenic bacteria and electrochemical magneto genosensing of the
double-tagged amplicon, Anal. Chem. 81 (14) (2009) 58125820.
[157] M.H.F. Meyer, M. Stehr, S. Bhuju, H.-J. Krause, M. Hartmanna, P. Miethe, M.
Singh, M. Keusgen, Magnetic biosensor for the detection of Yersinia pestis, J.
Microbiol. Methods 68 (2007) 218224.
[158] C. Palmer, J. Ye, A.G. Rand, S.V. Letcher, C.M. Mello, (100A-25) Detection of Vibrio
parahaemolyticus with a portable immunomagnetic focusing biosensor, Session
[159]
[160]
[161]
[162]
[163]
[164]
[165]
[166]
[167]
[168]
100A, Food Microbiol: General II, Ann. Meeting and Food Expo - Anaheim, California,
2002.
H. Lee, E. Sun, D. Ham, R. Weissleder, Chip-NMR biosensor for detection and molecular analysis of cells, Nat. Med. 14 (2008) 869874.
Y. Cheng, Y. Liu, J. Huang, K. Li, W. Zhang, Y. Xian, L. Jin, Combining biofunctional
magnetic nanoparticles and ATP bioluminescence for rapid detection of Escherichia
coli, Talanta 77 (2009) 13321336.
J.J. Pancrazio, J.P. Whelan, D.A. Borkholder, W. Ma, D.A. Stenger, Development
and application of cell based biosensors, Ann. Rev. Biomed. Eng. 27 (6) (1999)
697711.
T.H. Rider, M.S. Petrovick, F.E. Nargi, J.D. Harper, E.D. Schwoebel, R.H. Mathews, D.J.
Blanchard, L.T. Bortolin, A.M. Young, J. Chen, M.A. Hollis, A B-cell-based sensor for
rapid identication of pathogens, Science 301 (5630) (2003) 213215.
F.J. Picard, M.G. Bergeron, Rapid molecular theranostics in infectious diseases,
Drug Discov. Today 21 (7) (Nov. 2002).
N. Duran, P.D. Marcato, R. De Conti, O.L. Alves, F.T.M. Costa, M. Brocchi, Potential
use of silver nanoparticles on pathogenic bacteria, their toxicity and possible
mechanisms of action, J. Braz. Chem. Soc. 6 (21) (2010) 949959.
J.R. Morones, J.L. Elechiguerra, A. Camacho, K. Holt, J.B. Kouri, J.T. Ramirez, M.J.
Yacaman, The bactericidal effect of silver nanoparticles, Nanotechnology 16
(2005) 2346.
I. Sondi, B. Salopek-Sondi, Silver nanoparticles as antimicrobial agent: a case
study on E. coli as a model for Gram-negative bacteria, J. Colloid Interface Sci.
275 (2004) 177182.
J.S. Kim, E. Kuk, K.N. Yu, J.-H. Kim, S.J. Park, H.J. Lee, S.H. Kim, Y.K. Park, Y.H. Park,
C.-Y. Hwang, Y.-K. Kim, Y.-S. Lee, D.H. Jeong, M.-H. Cho, Antimicrobial effects of
silver nanoparticles, Nanomedicine 3 (2007) 95101.
J. Fabrega, C.J. Renshaw, R. Jamie, Lead interactions of silver nanoparticles with
Pseudomonas putida biolms, Environ. Sci. Technol. 43 (23) (2009) 90049009.
Potrebbero piacerti anche
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeDa EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeValutazione: 4 su 5 stelle4/5 (5794)
- Shoe Dog: A Memoir by the Creator of NikeDa EverandShoe Dog: A Memoir by the Creator of NikeValutazione: 4.5 su 5 stelle4.5/5 (537)
- The Yellow House: A Memoir (2019 National Book Award Winner)Da EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Valutazione: 4 su 5 stelle4/5 (98)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceDa EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceValutazione: 4 su 5 stelle4/5 (895)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersDa EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersValutazione: 4.5 su 5 stelle4.5/5 (344)
- The Little Book of Hygge: Danish Secrets to Happy LivingDa EverandThe Little Book of Hygge: Danish Secrets to Happy LivingValutazione: 3.5 su 5 stelle3.5/5 (399)
- Grit: The Power of Passion and PerseveranceDa EverandGrit: The Power of Passion and PerseveranceValutazione: 4 su 5 stelle4/5 (588)
- The Emperor of All Maladies: A Biography of CancerDa EverandThe Emperor of All Maladies: A Biography of CancerValutazione: 4.5 su 5 stelle4.5/5 (271)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaDa EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaValutazione: 4.5 su 5 stelle4.5/5 (266)
- Never Split the Difference: Negotiating As If Your Life Depended On ItDa EverandNever Split the Difference: Negotiating As If Your Life Depended On ItValutazione: 4.5 su 5 stelle4.5/5 (838)
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryDa EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryValutazione: 3.5 su 5 stelle3.5/5 (231)
- On Fire: The (Burning) Case for a Green New DealDa EverandOn Fire: The (Burning) Case for a Green New DealValutazione: 4 su 5 stelle4/5 (73)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureDa EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureValutazione: 4.5 su 5 stelle4.5/5 (474)
- Team of Rivals: The Political Genius of Abraham LincolnDa EverandTeam of Rivals: The Political Genius of Abraham LincolnValutazione: 4.5 su 5 stelle4.5/5 (234)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyDa EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyValutazione: 3.5 su 5 stelle3.5/5 (2259)
- The Unwinding: An Inner History of the New AmericaDa EverandThe Unwinding: An Inner History of the New AmericaValutazione: 4 su 5 stelle4/5 (45)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreDa EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreValutazione: 4 su 5 stelle4/5 (1090)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)Da EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Valutazione: 4.5 su 5 stelle4.5/5 (120)
- Her Body and Other Parties: StoriesDa EverandHer Body and Other Parties: StoriesValutazione: 4 su 5 stelle4/5 (821)
- Statistics Notes in The British Medical Journal (Bland JM, Altman DG. - NEJ)Documento95 pagineStatistics Notes in The British Medical Journal (Bland JM, Altman DG. - NEJ)pegazus_arNessuna valutazione finora
- Persistent Prelethal Stress Leads To Cellular AdaptationDocumento14 paginePersistent Prelethal Stress Leads To Cellular AdaptationMalliga SundareshanNessuna valutazione finora
- Perioperative Fasting and Feeding in Adults, Obstetric, Paediatric and Bariatric Population-Practice Guidelines From The Indian Society of AnaesthesiologistsDocumento29 paginePerioperative Fasting and Feeding in Adults, Obstetric, Paediatric and Bariatric Population-Practice Guidelines From The Indian Society of Anaesthesiologistsambitiousamit1Nessuna valutazione finora
- Cataract Surgery ProtocolsDocumento17 pagineCataract Surgery ProtocolsRahul ShastriNessuna valutazione finora
- Semmelweis' Handwashing & Lister's Antiseptic TechniqueDocumento22 pagineSemmelweis' Handwashing & Lister's Antiseptic TechniqueNewNessuna valutazione finora
- Presentation 1Documento12 paginePresentation 1Rizka FarahinNessuna valutazione finora
- CP 01708016Documento11 pagineCP 01708016telur_kudaNessuna valutazione finora
- Occupational HealthDocumento8 pagineOccupational HealthFemale calm0% (1)
- A Proposed Pediatric Medical Center Research ThesisDocumento3 pagineA Proposed Pediatric Medical Center Research ThesisMarielle AbanteNessuna valutazione finora
- Regulation of Iron MetabolismDocumento21 pagineRegulation of Iron MetabolismElita Maritan SNessuna valutazione finora
- Adoption Foster ApplicationDocumento4 pagineAdoption Foster Applicationlb8757Nessuna valutazione finora
- Magic of The Minimum Dose PDFDocumento221 pagineMagic of The Minimum Dose PDFminunat100% (1)
- Perkutan Kateter Vena Sentral Dibandingkan Perifer Kanula Untuk Pengiriman Nutrisi Parenteral Ada NeonatusDocumento3 paginePerkutan Kateter Vena Sentral Dibandingkan Perifer Kanula Untuk Pengiriman Nutrisi Parenteral Ada NeonatusmuslihudinNessuna valutazione finora
- Transient Tachypnea of The Newborn (TTN)Documento6 pagineTransient Tachypnea of The Newborn (TTN)Wivan Havilian DjohanNessuna valutazione finora
- NRC Managing Fever in Older Adults NSPDocumento5 pagineNRC Managing Fever in Older Adults NSPAulya FahnidaNessuna valutazione finora
- The Endocrine System and Feedback MechanismsDocumento8 pagineThe Endocrine System and Feedback MechanismsJame SmithNessuna valutazione finora
- Common Questions About Oppositional Defiant Disorder - American Family PhysicianDocumento12 pagineCommon Questions About Oppositional Defiant Disorder - American Family Physiciando lee100% (1)
- Brief Notes Parish of St. RafaelDocumento3 pagineBrief Notes Parish of St. RafaelAl F. Dela CruzNessuna valutazione finora
- Knowledge Regarding Immunization Among Mothers of Under Five ChildrenDocumento3 pagineKnowledge Regarding Immunization Among Mothers of Under Five ChildrenEditor IJTSRDNessuna valutazione finora
- Low-Dose Naltrexone (LDN) Fact Sheet 2016: Contact: Linda Elsegood Email: Skype Phone No'sDocumento8 pagineLow-Dose Naltrexone (LDN) Fact Sheet 2016: Contact: Linda Elsegood Email: Skype Phone No'scristiNessuna valutazione finora
- Rufenal EmulgelDocumento2 pagineRufenal EmulgelDr.2020Nessuna valutazione finora
- Full Download Test Bank For Essentials of Genetics 8th Edition by Klug PDF Full ChapterDocumento36 pagineFull Download Test Bank For Essentials of Genetics 8th Edition by Klug PDF Full Chapterfencingvesper9dgb04100% (17)
- HPP Neuro Paper GraserDocumento12 pagineHPP Neuro Paper GraserCaro ErazoNessuna valutazione finora
- Khalil, 2014Documento6 pagineKhalil, 2014Ibro DanNessuna valutazione finora
- Chapter - 2 Microorganisms (Continuation)Documento2 pagineChapter - 2 Microorganisms (Continuation)ARSHAD JAMILNessuna valutazione finora
- Cardiovascular Risk Factors in Airline PilotsDocumento4 pagineCardiovascular Risk Factors in Airline Pilotsluis11256Nessuna valutazione finora
- Spontaneous Pneumothorax - Management Feb15Documento12 pagineSpontaneous Pneumothorax - Management Feb15samuelNessuna valutazione finora
- Belgian MalinoisDocumento9 pagineBelgian MalinoislordnrNessuna valutazione finora
- Anatomy and Physiology of Male Reproductive SystemDocumento8 pagineAnatomy and Physiology of Male Reproductive SystemAdor AbuanNessuna valutazione finora
- Practice Test 05 - Hints & Solutions - Lakshya NEET 2024Documento15 paginePractice Test 05 - Hints & Solutions - Lakshya NEET 2024Pandey 14Nessuna valutazione finora